 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MELO3C006435P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 227aa MW: 25503.8 Da PI: 5.7815 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 33.2 | 9.2e-11 | 75 | 122 | 4 | 54 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksL 54
++ rEr RR+++N++f L l ++ +++K +K +iL A++ +++L
MELO3C006435P1 75 SKACRERLRREKLNDRFLDLSIALEPS---RHTKTNKPAILDDAIRVLNQL 122
5678***********************...******************999 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF47459 | 2.62E-12 | 68 | 139 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 11.985 | 71 | 122 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 6.61E-7 | 74 | 127 | No hit | No description |
| Pfam | PF00010 | 8.9E-8 | 75 | 122 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 2.3E-11 | 77 | 144 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 6.7E-8 | 77 | 128 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 227 aa Download sequence Send to blast |
MDPLEDSSCW DFLDCSFFPD ADNLINPSLD PLWPPNDSRK GSLADTDVSA GRVESEENDC 60 SRKRTRDGSC AGPSSKACRE RLRREKLNDR FLDLSIALEP SRHTKTNKPA ILDDAIRVLN 120 QLKNEAEDLK QTNEKLREEL KNLKAEKNDL RKEKIILKEE KEKMEQQLKS IAIPSPGLIP 180 GHPATYHAAS GKMAVFPGYG LIPMWQYLPP SIRDTSQDHE LRPPAA* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00102 | PBM | Transfer from AT4G14410 | Download |
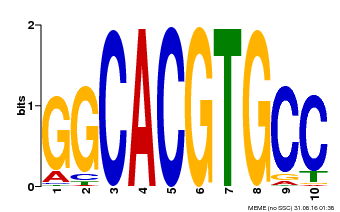
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681848 | 1e-137 | LN681848.1 Cucumis melo genomic scaffold, anchoredscaffold00006. | |||
| GenBank | LN713260 | 1e-137 | LN713260.1 Cucumis melo genomic chromosome, chr_6. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008438540.1 | 1e-166 | PREDICTED: transcription factor bHLH104-like | ||||
| Swissprot | Q8L467 | 4e-75 | BH104_ARATH; Transcription factor bHLH104 | ||||
| TrEMBL | A0A1S3AXA4 | 1e-164 | A0A1S3AXA4_CUCME; transcription factor bHLH104-like | ||||
| STRING | XP_008438540.1 | 1e-165 | (Cucumis melo) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF8179 | 25 | 37 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G14410.2 | 3e-66 | bHLH family protein | ||||



