 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MELO3C004728P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 491aa MW: 54798.8 Da PI: 6.9614 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 34.6 | 4.2e-11 | 184 | 225 | 4 | 45 |
XCHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 4 lkrerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaL 45
k rr+++NReAAr+sR+RKka+i++Le +L++ ++L
MELO3C004728P1 184 AKTLRRLAQNREAARKSRLRKKAYIQQLESSRIKLSQLEQDL 225
6889*************************9777777666655 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.20.5.170 | 1.1E-7 | 177 | 226 | No hit | No description |
| SMART | SM00338 | 1.1E-7 | 181 | 247 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 9.071 | 183 | 227 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF00170 | 3.0E-8 | 184 | 225 | IPR004827 | Basic-leucine zipper domain |
| CDD | cd14708 | 5.63E-22 | 185 | 236 | No hit | No description |
| SuperFamily | SSF57959 | 8.84E-7 | 185 | 226 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 188 | 203 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF14144 | 2.6E-31 | 268 | 342 | IPR025422 | Transcription factor TGA like domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 491 aa Download sequence Send to blast |
MANQHSATLP NFHGIIHHNP SLPFINPEGS AFDFGELEEA IVLQGVKLGN DEPKSPNFLT 60 GRPAATLEMF PSWPIRFQQT PTLGGGSKSE STDSGSANIN NTLTSKIELE MESGSPINRR 120 TCSSNQGLFD QNHHHHHHLL HLQHLQSEFE DDALRTEASS QQNQSPLKEK RKGGGSTSER 180 QLDAKTLRRL AQNREAARKS RLRKKAYIQQ LESSRIKLSQ LEQDLHRARS QGLFVGACGG 240 VMGGNISSGA AIFDMEYARW LDEDHRLMAE LRAALQGHLP DGDLRAIVDS YISHYDEIFH 300 LKGVAAKSDV FHLITGMWMT PAERCFLWIG GFRPSKLIEM LIPQIDTLTE QQAMGICNLQ 360 RSSQETEDAL YQGLEQLQHS LIITIAGTAV VDGINHMALA AGKLSNLEGF IRQADMLRQQ 420 TLHQLHRILT VRQAARCFVV IGEYYGRLRA LSSLWVSRPR ESSCLNDESS CQTTTELQMI 480 QNSHTHFPNF * |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Together with TGA10, basic leucine-zipper transcription factor required for anther development, probably via the activation of SPL expression in anthers and via the regulation of genes with functions in early and middle tapetal development (PubMed:20805327). Required for signaling responses to pathogen-associated molecular patterns (PAMPs) such as flg22 that involves chloroplastic reactive oxygen species (ROS) production and subsequent expression of H(2)O(2)-responsive genes (PubMed:27717447). {ECO:0000269|PubMed:20805327, ECO:0000269|PubMed:27717447}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00131 | DAP | Transfer from AT1G08320 | Download |
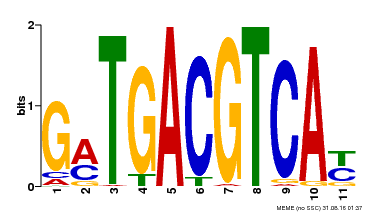
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by flg22 in leaves. {ECO:0000269|PubMed:27717447}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681928 | 1e-150 | LN681928.1 Cucumis melo genomic scaffold, anchoredscaffold00004. | |||
| GenBank | LN713266 | 1e-150 | LN713266.1 Cucumis melo genomic chromosome, chr_12. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008461530.1 | 0.0 | PREDICTED: transcription factor HBP-1b(c38)-like | ||||
| Swissprot | Q93XM6 | 0.0 | TGA9_ARATH; Transcription factor TGA9 | ||||
| TrEMBL | A0A1S3CEU5 | 0.0 | A0A1S3CEU5_CUCME; transcription factor HBP-1b(C38)-like | ||||
| STRING | XP_008461530.1 | 0.0 | (Cucumis melo) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF4555 | 33 | 57 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G08320.3 | 0.0 | bZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



