 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cla021184 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Citrullus
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 417aa MW: 44242.4 Da PI: 9.506 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 71.6 | 1.2e-22 | 281 | 343 | 1 | 63 |
XXXXCHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 1 ekelkrerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkkevaklksev 63
ekelkrerrkq+NRe+ArrsR+RK+ae+eeL++kv++L+aeN a ++e+++l+++++klk+e+
Cla021184 281 EKELKRERRKQSNRESARRSRLRKQAETEELARKVDSLTAENVAIRSEISRLSENSEKLKKEN 343
89***********************************************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF07777 | 4.9E-34 | 1 | 97 | IPR012900 | G-box binding protein, multifunctional mosaic region |
| Pfam | PF16596 | 6.2E-27 | 132 | 263 | No hit | No description |
| Gene3D | G3DSA:1.20.5.170 | 2.7E-16 | 275 | 341 | No hit | No description |
| SMART | SM00338 | 8.3E-21 | 281 | 345 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF00170 | 8.9E-21 | 281 | 343 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 13.231 | 283 | 346 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 3.38E-10 | 284 | 340 | No hit | No description |
| CDD | cd14702 | 9.73E-21 | 286 | 334 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 288 | 303 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0005829 | Cellular Component | cytosol | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 417 aa Download sequence Send to blast |
MGTSEEAKSV KTEKPSSPTP DQNSVPNSAS IHVFPDWAAM QAYYGPRVAV PPYYNSAVAS 60 GHAPHPYMWA PPQMIPPYGT PYAAIYSHGG VYAHPAVSMG PHSHGPGVPS SPGAAPPLSI 120 ETPSKVSGNS SQGLMKKLKG FDGLAMSIGN VSTETAEGGA EHGQSESMET EGSSDGSDGT 180 TAGANQTKRK RSREGTPTTG KDAKIEPQAS PVTAAEMNES SSKLLGTTKA ANATGKLGSV 240 ISPGMSTALE LRNPSSINAM TSPTTVPPCS VLPSEVWLQN EKELKRERRK QSNRESARRS 300 RLRKQAETEE LARKVDSLTA ENVAIRSEIS RLSENSEKLK KENSTLMEKL KSAQSGRTEA 360 LDMNEKRMQQ PVSTETKGPV NKSINEESII CKKNSSSGAK LRQLLDTSPR ADAVAAS |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 297 | 303 | RRSRLRK |
| 2 | 297 | 304 | RRSRLRKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds to the G-box-like motif (5'-ACGTGGC-3') of the chalcone synthase (CHS) gene promoter. G-box and G-box-like motifs are defined in promoters of certain plant genes which are regulated by such diverse stimuli as light-induction or hormone control. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00318 | DAP | Transfer from AT2G46270 | Download |
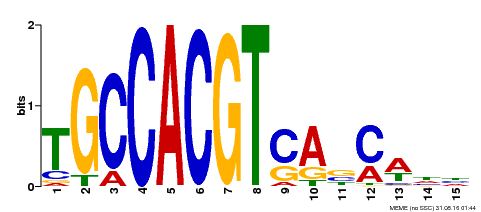
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By light. | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681848 | 1e-95 | LN681848.1 Cucumis melo genomic scaffold, anchoredscaffold00006. | |||
| GenBank | LN713260 | 1e-95 | LN713260.1 Cucumis melo genomic chromosome, chr_6. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008437949.1 | 0.0 | PREDICTED: common plant regulatory factor 1 isoform X2 | ||||
| Swissprot | Q99089 | 1e-125 | CPRF1_PETCR; Common plant regulatory factor 1 | ||||
| TrEMBL | A0A1S3AV69 | 0.0 | A0A1S3AV69_CUCME; common plant regulatory factor 1 isoform X2 | ||||
| TrEMBL | E5GCG9 | 0.0 | E5GCG9_CUCME; Bzip transcription factor | ||||
| STRING | XP_008437947.1 | 0.0 | (Cucumis melo) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF5301 | 34 | 55 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46270.1 | 1e-110 | G-box binding factor 3 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



