 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cla020642 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Citrullus
|
||||||||
| Family | WRKY | ||||||||
| Protein Properties | Length: 380aa MW: 41065.3 Da PI: 10.0979 | ||||||||
| Description | WRKY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | WRKY | 97 | 1.3e-30 | 309 | 366 | 3 | 59 |
-SS-EEEEEEE--TT-SS-EEEEEE-ST.T---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 3 DgynWrKYGqKevkgsefprsYYrCtsa.gCpvkkkversaedpkvveitYegeHnhe 59
D y+WrKYGqK++kgs++pr+YY+C+s gCp++k+ver+ +dp+++++tYe++Hnh
Cla020642 309 DDYSWRKYGQKPIKGSPHPRGYYKCSSLrGCPARKHVERALDDPTMLIVTYENDHNHA 366
99************************998****************************6 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF10533 | 3.5E-18 | 260 | 305 | IPR018872 | Zn-cluster domain |
| Gene3D | G3DSA:2.20.25.80 | 1.8E-32 | 296 | 366 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 29.585 | 302 | 368 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 3.4E-26 | 305 | 366 | IPR003657 | WRKY domain |
| SMART | SM00774 | 3.1E-36 | 307 | 367 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 3.5E-26 | 309 | 365 | IPR003657 | WRKY domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 380 aa Download sequence Send to blast |
MAVELMVGFG DATTPTNFTP KMEENAAVSA VQEAASAGIQ SVQNFLRLMS HTPNHQHSQD 60 DSSASTTPNA IAGYEAVADS VVSKFKKVIS LLDRSRTGHA RFRRAPVLTT TPPPPPPPKV 120 KPQHQDPSSS SPISVPQIQV KKQESVSAFK VYCPTPSPVV RLPPLPHNNS HQPSHPPNTF 180 QAQQNTSSVV LKNGSVDRKD ATTTINFAAS PPISAANSYI SSLTGDTESL QPSLSSGFQF 240 THMSQVSSAG KPPLSSSSLK RKCNSMEDSA MKCGSSSGRC HCSKKRKNRI KRVIRVPAVS 300 SKLADIPPDD YSWRKYGQKP IKGSPHPRGY YKCSSLRGCP ARKHVERALD DPTMLIVTYE 360 NDHNHAHSTE TPAALVLESS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2ayd_A | 2e-20 | 295 | 365 | 2 | 71 | WRKY transcription factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. Interacts specifically with the W box (5'-(T)TGAC[CT]-3'), a frequently occurring elicitor-responsive cis-acting element (By similarity). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00450 | DAP | Transfer from AT4G24240 | Download |
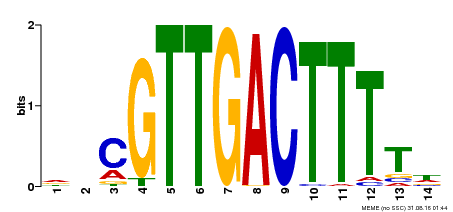
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salicylic acid and strongly during leaf senescence. {ECO:0000269|PubMed:11449049, ECO:0000269|PubMed:11722756}. | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | GU984027 | 0.0 | GU984027.1 Cucumis sativus WRKY protein (WRKY33) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_022937239.1 | 0.0 | probable WRKY transcription factor 7 isoform X2 | ||||
| Refseq | XP_023536442.1 | 0.0 | probable WRKY transcription factor 7 isoform X2 | ||||
| Swissprot | Q9STX0 | 1e-108 | WRKY7_ARATH; Probable WRKY transcription factor 7 | ||||
| TrEMBL | A0A1S3BVC4 | 0.0 | A0A1S3BVC4_CUCME; probable WRKY transcription factor 7 isoform X2 | ||||
| STRING | XP_008452579.1 | 0.0 | (Cucumis melo) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2177 | 34 | 86 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G24240.1 | 4e-77 | WRKY DNA-binding protein 7 | ||||



