 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cla020369 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Citrullus
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 357aa MW: 39026.5 Da PI: 8.0463 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 104.6 | 5.6e-33 | 206 | 261 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W++eLH+rFv+a++qLGG+++AtPk+i+elmkv+gLt+++vkSHLQkYRl+
Cla020369 206 KQRRCWSSELHRRFVHALQQLGGPHVATPKQIRELMKVDGLTNDEVKSHLQKYRLH 261
79****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 6.2E-29 | 203 | 264 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 3.05E-18 | 203 | 263 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 14.807 | 203 | 263 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 1.1E-27 | 206 | 261 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 6.4E-7 | 208 | 259 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 357 aa Download sequence Send to blast |
MDFLHNFLDI RVGMDLRHYA DALDQERRKI QVFQRELPLC LELVTRAIES CRQQLSGVST 60 DSSEHTSSDG PVLEEFIPIN KPSVNSHFEI EEDEQSKPEK IELGRSDWLK SAQLWNQSPD 120 PPVTEDVAKE TTSSVVEVKK NVGGGAFQPF QKEKTTPLKT ESVVVGKSDG SSPIPEATTS 180 STAETASKGG GGTSKREDKE SQTQRKQRRC WSSELHRRFV HALQQLGGPH VATPKQIREL 240 MKVDGLTNDE VKSHLQKYRL HARRPTNSAM QDSGSSSAPQ QFVVVGSIWV PPPPPPKYTV 300 ATAAKARNGI YAPVATAVRR QTTAEEGKAS HSGGVVHSNS PATSSCTHTT TTSPSAA |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in phosphate signaling in roots. {ECO:0000250|UniProtKB:Q9FX67}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00160 | DAP | Transfer from AT1G25550 | Download |
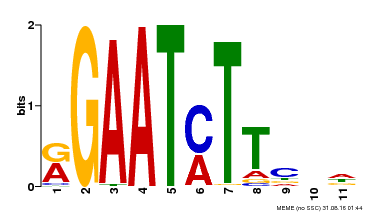
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681896 | 1e-115 | LN681896.1 Cucumis melo genomic scaffold, anchoredscaffold00016. | |||
| GenBank | LN713264 | 1e-115 | LN713264.1 Cucumis melo genomic chromosome, chr_10. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008446559.1 | 0.0 | PREDICTED: LOW QUALITY PROTEIN: transcription factor LUX | ||||
| Swissprot | Q9FPE8 | 2e-89 | HHO3_ARATH; Transcription factor HHO3 | ||||
| TrEMBL | A0A1S3BG56 | 0.0 | A0A1S3BG56_CUCME; LOW QUALITY PROTEIN: transcription factor LUX | ||||
| STRING | XP_008446559.1 | 0.0 | (Cucumis melo) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3918 | 33 | 63 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G68670.1 | 2e-86 | G2-like family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



