 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cla017518 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Citrullus
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 370aa MW: 40084.3 Da PI: 9.5383 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 39.5 | 9.9e-13 | 298 | 343 | 6 | 55 |
HHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 6 nerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
+++Er RR ri +++ +L++l+P+ +k + a++L +Av+YIk+Lq
Cla017518 298 SIAERVRRTRISERMRKLQDLVPNM----DKQTNTADMLDLAVDYIKELQ 343
689*********************7....688*****************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50888 | 15.32 | 292 | 342 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 6.67E-16 | 295 | 352 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 1.85E-12 | 296 | 345 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 5.9E-15 | 297 | 353 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 4.0E-12 | 298 | 348 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 1.2E-9 | 298 | 343 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 370 aa Download sequence Send to blast |
MGSNTHQSFQ QPNSALLRFR SAPSSLFADF AQGIDSKRSN PFEGSESERL VSRFGNRGGG 60 GNSNDSESPA AGNYSSGLPP HYPRLSSAVN CSSSSSTTTS SSSSSSSSSS MCSSLGFLGP 120 NLVRQSSSPA GVFSQLNQNG YGGGSFSRLS VNNNGGEVSP SSNRLNSQIS FSSLLPSSLG 180 MFPQISEQVV GNEKLVNSNN GETQFFTPSG FPFASWNESS QFSETFPDIK REPDSNKKLF 240 SSSHQNGEIG NRVHLLSHHL SLPKNASDVA SIEKLLQLQD AVPCRIRAKR GCATHPRSIA 300 ERVRRTRISE RMRKLQDLVP NMDKQTNTAD MLDLAVDYIK ELQKQFKTLS DNRANCVCLN 360 MQKSVSNQIM |
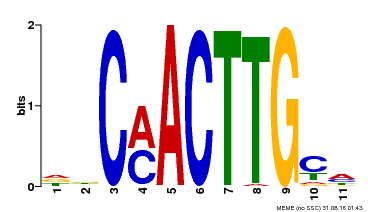
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00308 | DAP | Transfer from AT2G42280 | Download |
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681841 | 1e-160 | LN681841.1 Cucumis melo genomic scaffold, anchoredscaffold00011. | |||
| GenBank | LN713258 | 1e-160 | LN713258.1 Cucumis melo genomic chromosome, chr_4. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004136581.1 | 0.0 | PREDICTED: transcription factor bHLH130 | ||||
| Swissprot | Q66GR3 | 7e-93 | BH130_ARATH; Transcription factor bHLH130 | ||||
| TrEMBL | A0A0A0LH01 | 0.0 | A0A0A0LH01_CUCSA; Uncharacterized protein | ||||
| STRING | XP_004136581.1 | 0.0 | (Cucumis sativus) | ||||
| STRING | XP_004168456.1 | 0.0 | (Cucumis sativus) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF5177 | 32 | 53 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G42280.1 | 3e-89 | bHLH family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



