 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cla017323 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Citrullus
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 1009aa MW: 110808 Da PI: 6.3359 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 131.8 | 2.4e-41 | 155 | 232 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+C adls+ak+yhrrhkvCe+hska+++lv++++qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk ++
Cla017323 155 VCQVEDCGADLSNAKDYHRRHKVCETHSKASNALVANVMQRFCQQCSRFHVLQEFDEGKRSCRRRLAGHNKRRRKVNP 232
6**************************************************************************875 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 1.0E-33 | 149 | 217 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.035 | 153 | 230 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 3.66E-38 | 154 | 235 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 6.1E-30 | 156 | 229 | IPR004333 | Transcription factor, SBP-box |
| Gene3D | G3DSA:1.25.40.20 | 1.9E-7 | 748 | 907 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 4.14E-8 | 750 | 891 | No hit | No description |
| SuperFamily | SSF48403 | 2.64E-8 | 775 | 891 | IPR020683 | Ankyrin repeat-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1009 aa Download sequence Send to blast |
MEAGYGGEAC QLYGMGPMDL RAAVGKRNLE WDLNDWKWDG DLFIARPLNT VESDHLSRQL 60 FPIVSGIPLT NGGSSNSSSS CSDEANMGIE KGKREVEKRR RVTVIEDENL NDEARTLSLK 120 VGGNGSQIVE REAGSWEGTS GKKTKLAGGN SNRAVCQVED CGADLSNAKD YHRRHKVCET 180 HSKASNALVA NVMQRFCQQC SRFHVLQEFD EGKRSCRRRL AGHNKRRRKV NPDNVVNGNS 240 PPDEQTSSYL LLTLLRILAN LHSNGSNQTT DQDLLSHLIR SLACQSSEHG GKNLSGILHE 300 PQSLLNNGAL MGKSDLVSTF LSNGPQASLR SSKPHDTIPI SETPVQAIGR GGDTPAISSI 360 KPSTSNSPPA YSEIRDSTVG QCKMMNFDLN DAYVDSDDGM EDIERQTLPV HMGTSSLECP 420 SWVQQDSHQS SPPQTSGNSD SASAQSPSSS AGEAQSRTDR IILKLFGKAP NDFPHVLRAQ 480 VLDWLSHSPT EIESYIRPGC VVLTIYLRQT EASWDNLCHD LSTSFNRLLD VSDDAFWRTG 540 WVYVRVQHQI AIVYHGQVVV DTSLPLRNKN YCRITSVNPV AVSTSKKAMF SVKGINLTQP 600 TTRLLCAIEG KYLTQEASDG STESGEDDSR CVTFSCSIPV VYGRGFIEVE DDGFSSSSFP 660 FIVAEEDVCS EICTLQSALE LTETCSNSGE TAELEGRSNA MEFIHEIGWL FHRNQLKSRL 720 GHLDPNENLF SLPRFKWLME FSMDHDWCAV VKKLLDVLRD GNVDAGGHSS LNLALMEMGL 780 LHRAVRKNSR SLVELLLRYP SKVKDASSSE DSASIDGGAD SFLFRPNAIG PAGLTPLHIA 840 AGKDDSEDVL DALTNDPGMV GVEAWKSARD STGSTPEDYA RLRGHYSYIR LVQRKINKRS 900 AAGHVVLDIP SSLSDSSWNQ KPNTDLASSR FEIGRMELKP SQQHCKLCVR KPLGCGTPSS 960 ASLVYRPAML SMVAIAAVCV CVALLFKSSP EVLYVFRPFR WELLDYGTS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 8e-32 | 149 | 229 | 4 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 97 | 101 | KRRRV |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000269|PubMed:16554053}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
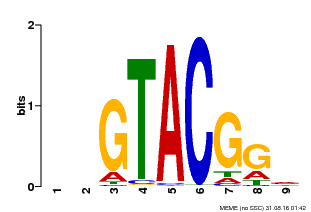
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681827 | 0.0 | LN681827.1 Cucumis melo genomic scaffold, anchoredscaffold00018. | |||
| GenBank | LN713258 | 0.0 | LN713258.1 Cucumis melo genomic chromosome, chr_4. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004148578.1 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 1 | ||||
| Swissprot | Q9S7P5 | 0.0 | SPL12_ARATH; Squamosa promoter-binding-like protein 12 | ||||
| TrEMBL | A0A0A0LEQ2 | 0.0 | A0A0A0LEQ2_CUCSA; Uncharacterized protein | ||||
| STRING | XP_008447446.1 | 0.0 | (Cucumis melo) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1539 | 34 | 95 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



