 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cla015673 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Citrullus
|
||||||||
| Family | WRKY | ||||||||
| Protein Properties | Length: 435aa MW: 47724.7 Da PI: 6.1695 | ||||||||
| Description | WRKY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | WRKY | 93 | 2.1e-29 | 110 | 167 | 2 | 60 |
--SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS-- CS
WRKY 2 dDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhek 60
+DgynWrKYGqK vkg+ f rsYYrCt+++C+vkk++er++ d+k+++++Y g+H+h+k
Cla015673 110 EDGYNWRKYGQKLVKGNVFVRSYYRCTHPTCMVKKQLERTH-DGKITDTVYFGQHDHPK 167
7***************************************9.***************85 PP
| |||||||
| 2 | WRKY | 106 | 1.9e-33 | 282 | 340 | 1 | 59 |
---SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 1 ldDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhe 59
++Dgy+WrKYGqK vkg+++prsYYrC+s+gCpvkk+ver++ dpkvv +tYeg+H+h+
Cla015673 282 VNDGYRWRKYGQKLVKGNPNPRSYYRCSSPGCPVKKHVERASYDPKVVLTTYEGQHDHD 340
58********************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.20.25.80 | 9.3E-26 | 97 | 167 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 21.551 | 104 | 168 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 6.67E-24 | 104 | 167 | IPR003657 | WRKY domain |
| SMART | SM00774 | 2.7E-32 | 109 | 167 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 8.6E-23 | 110 | 166 | IPR003657 | WRKY domain |
| Gene3D | G3DSA:2.20.25.80 | 4.3E-35 | 268 | 342 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 5.23E-29 | 274 | 342 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 35.84 | 277 | 342 | IPR003657 | WRKY domain |
| SMART | SM00774 | 6.0E-38 | 282 | 341 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 4.9E-26 | 283 | 339 | IPR003657 | WRKY domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009863 | Biological Process | salicylic acid mediated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 435 aa Download sequence Send to blast |
MVSSGDQLEN EVDSDQLEHE NSSESQPQTS QDDPGGTNAS KSDHKCTGAA SITLEEAVKQ 60 PEVTIEQVDR GEISNIVTEK VTHKPITAEQ NSHSDLKVCI TSTIREKVSE DGYNWRKYGQ 120 KLVKGNVFVR SYYRCTHPTC MVKKQLERTH DGKITDTVYF GQHDHPKAQP HIPVAVGVVT 180 MVEEKLDEHA SGNSQDKTSI APGQTPHQTE LADMRQPSSV MASDDVKDEA SKRSRINDEV 240 DSDDTPDLKR EKKRCNIDVT VADKSTVESR VVVQTPSEVD IVNDGYRWRK YGQKLVKGNP 300 NPRSYYRCSS PGCPVKKHVE RASYDPKVVL TTYEGQHDHD MPPTRTVTLN SVGSTAAHSD 360 ETKPKPVGSS IGHDTVVHAT KDSLNNSSSE GKLIEKNGKS NATEASDGIV LDMVVNPSPG 420 VASGQNKQLK VAIES |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2ayd_A | 6e-38 | 108 | 343 | 1 | 75 | WRKY transcription factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. Binds to a 5'-CGTTGACCGAG-3' consensus core sequence which contains a W box, a frequently occurring elicitor-responsive cis-acting element. {ECO:0000269|PubMed:8972846}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00094 | SELEX | Transfer from AT2G04880 | Download |
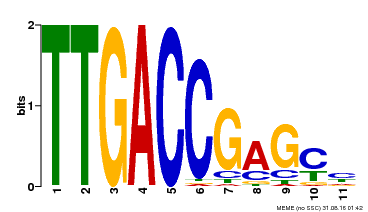
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salicylic acid (SA). {ECO:0000269|PubMed:17264121}. | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | GU984034 | 0.0 | GU984034.1 Cucumis sativus WRKY protein (WRKY49) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001267610.1 | 0.0 | probable WRKY transcription factor 20-like | ||||
| Refseq | XP_011658539.1 | 0.0 | PREDICTED: probable WRKY transcription factor 20-like isoform X1 | ||||
| Swissprot | Q9SI37 | 2e-87 | WRKY1_ARATH; WRKY transcription factor 1 | ||||
| TrEMBL | E7CEY3 | 0.0 | E7CEY3_CUCSA; WRKY protein | ||||
| STRING | XP_004144541.1 | 0.0 | (Cucumis sativus) | ||||
| STRING | XP_004164058.1 | 0.0 | (Cucumis sativus) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF9358 | 33 | 43 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G04880.2 | 1e-95 | zinc-dependent activator protein-1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



