 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cagra.1961s0066.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Capsella
|
||||||||
| Family | BES1 | ||||||||
| Protein Properties | Length: 334aa MW: 36232.2 Da PI: 9.3741 | ||||||||
| Description | BES1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | DUF822 | 207.4 | 4.1e-64 | 18 | 157 | 2 | 144 |
DUF822 2 gsgrkptwkErEnnkrRERrRRaiaakiyaGLRaqGnyklpkraDnneVlkALcreAGwvvedDGttyrkgskpleeaeaagssasaspe 91
+++rkp+w+ErEnn+rRERrRRa+aakiy+GLRaqGny+lpk++DnneVlkALc eAGwvve+DGttyrkg kpl ++agss++a+p
Cagra.1961s0066.1.p 18 ATRRKPSWRERENNRRRERRRRAVAAKIYTGLRAQGNYNLPKHCDNNEVLKALCSEAGWVVEEDGTTYRKGHKPL-PGDMAGSSSRATPY 106
689************************************************************************.************** PP
DUF822 92 sslqsslkssalaspvesysaspksssfpspssldsislasaasllpvlsvls 144
ss ++s+ ss++ sp+ sy+ sp+sssfpsps++ ++ ++++p+l++ +
Cagra.1961s0066.1.p 107 SSHNQSPLSSTFDSPILSYQVSPSSSSFPSPSRVGDPHNI--STIFPFLRNGG 157
*********************************8776655..67888887765 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF05687 | 2.9E-61 | 19 | 140 | IPR008540 | BES1/BZR1 plant transcription factor, N-terminal |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009742 | Biological Process | brassinosteroid mediated signaling pathway | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005829 | Cellular Component | cytosol | ||||
| GO:0001046 | Molecular Function | core promoter sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 334 aa Download sequence Send to blast |
MTSDGATSTS AAAAAAMATR RKPSWREREN NRRRERRRRA VAAKIYTGLR AQGNYNLPKH 60 CDNNEVLKAL CSEAGWVVEE DGTTYRKGHK PLPGDMAGSS SRATPYSSHN QSPLSSTFDS 120 PILSYQVSPS SSSFPSPSRV GDPHNISTIF PFLRNGGIPS SLPPLRISNS CPVTPPVSSP 180 TSRNPKPLPT WESFTKQSMS MAAKQSMTSL NYPFYAVSAP ASPTHHRQFH APATIPECDE 240 SDSSTVDSGH WISFQKFSQQ PFHGASVVPA SPTFNLVKPA PQQLSPNTAA TQEIGQSSDF 300 KFENSQVKPW EGERIHDVAM EDLELTLGNG KSS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5zd4_A | 4e-36 | 20 | 102 | 371 | 453 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| 5zd4_B | 4e-36 | 20 | 102 | 371 | 453 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| 5zd4_C | 4e-36 | 20 | 102 | 371 | 453 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| 5zd4_D | 4e-36 | 20 | 102 | 371 | 453 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 19 | 38 | RRKPSWRERENNRRRERRRR |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Positive regulator of brassinosteroid (BR) signaling. Transcription factor that activates target gene expression by binding specifically to the DNA sequence 5'-CANNTG-3'(E box) through its N-terminal domain. Can bind individually to the promoter as a homodimer or synergistically as a heterodimer with BIM1, BIM2 or BIM3. The C-terminal domain is probably involved in transcriptional activation (PubMed:12007405, PubMed:15680330, PubMed:18467490, PubMed:19170933). Recruits the transcription elongation factor IWS1 to control BR-regulated gene expression (PubMed:20139304). Forms a trimeric complex with IWS1 and ASHH2/SDG8 to regulate BR-regulated gene expression (PubMed:24838002). Promotes quiescent center (QC) self-renewal by cell divisions in the primary root. Binds to the E-boxes of the BRAVO promoter to repress its expression (PubMed:24981610). {ECO:0000269|PubMed:12007405, ECO:0000269|PubMed:15680330, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:19170933, ECO:0000269|PubMed:20139304, ECO:0000269|PubMed:24838002, ECO:0000269|PubMed:24981610}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00073 | ChIP-chip | Transfer from AT1G19350 | Download |
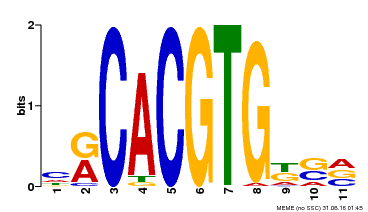
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Cagra.1961s0066.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY065041 | 0.0 | AY065041.1 Arabidopsis thaliana At1g19350/F18O14_4 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006305153.2 | 0.0 | protein BRASSINAZOLE-RESISTANT 2 | ||||
| Refseq | XP_023644976.1 | 0.0 | protein BRASSINAZOLE-RESISTANT 2 | ||||
| Swissprot | Q9LN63 | 0.0 | BZR2_ARATH; Protein BRASSINAZOLE-RESISTANT 2 | ||||
| TrEMBL | R0GQL6 | 0.0 | R0GQL6_9BRAS; Uncharacterized protein (Fragment) | ||||
| STRING | XP_006305153.1 | 0.0 | (Capsella rubella) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2472 | 27 | 72 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G19350.3 | 0.0 | BES1 family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Cagra.1961s0066.1.p |



