 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cagra.1226s0039.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Capsella
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 349aa MW: 38153.8 Da PI: 8.3148 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 59.8 | 6.5e-19 | 119 | 169 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
++++GVr+++ +g+W+AeIrdp + r r +lg++ taeeAa +++a+ +l+g
Cagra.1226s0039.1.p 119 KKFRGVRQRP-WGKWAAEIRDPLK---RVRLWLGTYNTAEEAAMVYDNAAIQLRG 169
59********.**********754...5*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 4.45E-32 | 119 | 176 | No hit | No description |
| SuperFamily | SSF54171 | 9.15E-22 | 120 | 178 | IPR016177 | DNA-binding domain |
| PROSITE profile | PS51032 | 23.406 | 120 | 177 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 1.1E-29 | 120 | 177 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 3.9E-37 | 120 | 183 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 6.9E-12 | 120 | 169 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 7.9E-10 | 121 | 132 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 7.9E-10 | 143 | 159 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0042991 | Biological Process | transcription factor import into nucleus | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:0048825 | Biological Process | cotyledon development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042802 | Molecular Function | identical protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 349 aa Download sequence Send to blast |
MEAEKKMVVP RIKFTEHKTN TTRIVPDLTN THQTRVLRIS VTDPDATDSS SDDEDEARNR 60 FVSKRRRVKK FVNEVVLDSR AVSGSCSQIE SKKSRKRAVK SESTASQVVS ATTATTGEKK 120 FRGVRQRPWG KWAAEIRDPL KRVRLWLGTY NTAEEAAMVY DNAAIQLRGP DALTNFSVNP 180 TTTTKTEKKS PPPSPVKKKK KKNNKIKKSV TASSSVSGSS SNDCLCSPVS VLRSPFTVDD 240 FSGSSSPAAA AVVVKEEPSM TTVSEAFSDF SAPLFSDDDL FDFRSSVVPD YLGLGGGDLF 300 GEDIFTAEMC TDMNFGFDFG SSGLSSWHME DHFQDIGDLF GSDPLLAV* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1gcc_A | 4e-20 | 119 | 176 | 1 | 59 | ETHYLENE RESPONSIVE ELEMENT BINDING FACTOR 1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 63 | 67 | KRRRV |
| 2 | 197 | 207 | KKKKKNNKIKK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Component of the cytokinin signaling pathway involved in cotyledons, leaves, and embryos development. Probably acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. May be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways (By similarity). {ECO:0000250, ECO:0000269|PubMed:16832061}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00027 | PBM | Transfer from AT4G23750 | Download |
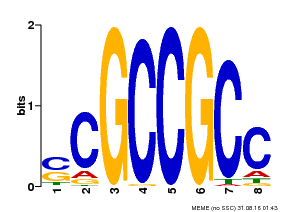
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Cagra.1226s0039.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By cytokinins. {ECO:0000269|PubMed:16832061}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006284039.1 | 0.0 | ethylene-responsive transcription factor CRF2 | ||||
| Swissprot | Q9SUQ2 | 0.0 | CRF2_ARATH; Ethylene-responsive transcription factor CRF2 | ||||
| TrEMBL | R0F592 | 0.0 | R0F592_9BRAS; Uncharacterized protein | ||||
| STRING | Cagra.1226s0039.1.p | 0.0 | (Capsella grandiflora) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM10 | 28 | 1650 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G23750.2 | 1e-112 | cytokinin response factor 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Cagra.1226s0039.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



