 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cagra.0248s0103.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Capsella
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1013aa MW: 113094 Da PI: 6.5394 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 188.3 | 7.3e-59 | 19 | 136 | 2 | 118 |
CG-1 2 lke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahsee 90
l+e ++rwl++ ei++iL n++k+++++e+++rp+sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+hekLKvg+++vl+cyYah+e+
Cagra.0248s0103.1.p 19 LSEaQHRWLRPAEICEILRNHQKFHIASEPPNRPPSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHEKLKVGSIDVLHCYYAHGED 108
45559************************************************************************************* PP
CG-1 91 nptfqrrcywlLeeelekivlvhylevk 118
n++fqrrcyw+Le++l +iv+vhylevk
Cagra.0248s0103.1.p 109 NENFQRRCYWMLEQDLMHIVFVHYLEVK 136
*************************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 86.275 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 3.6E-87 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 8.0E-52 | 21 | 135 | IPR005559 | CG-1 DNA-binding domain |
| SuperFamily | SSF81296 | 3.85E-13 | 422 | 506 | IPR014756 | Immunoglobulin E-set |
| SuperFamily | SSF48403 | 6.53E-19 | 603 | 714 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 19.28 | 603 | 714 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 6.4E-18 | 604 | 715 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 2.3E-7 | 608 | 680 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 2.06E-14 | 609 | 712 | No hit | No description |
| PROSITE profile | PS50088 | 8.71 | 620 | 652 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 3200 | 620 | 649 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 10.873 | 653 | 685 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.0029 | 653 | 682 | IPR002110 | Ankyrin repeat |
| SMART | SM00015 | 0.32 | 828 | 850 | IPR000048 | IQ motif, EF-hand binding site |
| SuperFamily | SSF52540 | 7.97E-8 | 829 | 878 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| PROSITE profile | PS50096 | 7.858 | 830 | 858 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0019 | 830 | 849 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0092 | 851 | 873 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.304 | 852 | 876 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 4.4E-4 | 854 | 873 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1013 aa Download sequence Send to blast |
MADRGSFGFA PRLDIKQLLS EAQHRWLRPA EICEILRNHQ KFHIASEPPN RPPSGSLFLF 60 DRKVLRYFRK DGHNWRKKKD GKTVKEAHEK LKVGSIDVLH CYYAHGEDNE NFQRRCYWML 120 EQDLMHIVFV HYLEVKGNRM STGGTKENHS NSLSGTGSVN VDSTATRSSI LSPLCEDADS 180 GDSHQASSSL QPNPELQIMH HQNASTMNSF NTSSVLGNRD GWTSAPAIGI VPHVHGNRVK 240 ESDSQRSGSL LTAEHLRNPL QNQVNWQIPV QESLPLQKWP MDSHSCMTDA TDLALFGQRP 300 HGNFGTFSSL LGCQNQQPSS FQAPFTNNEA AYIPKLGPED LIYEASANQT LPLRKALLKT 360 EDSLKKVDSF SRWVSKELGE MEDLQMQSSS GGIAWTSVEC ETAAAGSSLS PSLSEVQRFT 420 MIDFWPKWTQ TDSEVEVMVI GTFLLSPKEV TSYSWSCMFG EVEVPAEILV DGVLCCHAPP 480 HEVGRVPFYI TCSDRFSCSE VREFDFLPGS TRKLNATDVY GANTIEASLH LRFENLLALR 540 SSVQEHHIFE NVGQKRRKIS KIMLLMDEKE SLLPGTIEKD STELEAKERL IREEFEDKLY 600 LWLIHKVTEE GKGPNILDEV GQGVLHLAAA LGYDWAIKPI LAAGVSINFR DANGWSALHW 660 AAFSGREDTV AVLVSLGADA GALADPSPEH PLGKTAADLA YGNGHRGISG FLAESSLTSY 720 LEKLTVDAKE NSSADSSGAK AVLTVAERTA TPMSYGDVPE TLSMKDSLTA VLNATQAADR 780 LHQVFRMQSF QRKQLSELGG DNEFDISDEL AVSFAAAKTK KPGHSNGAVH AAAVQIQKKY 840 RGWKKRKEFL LIRQRIVKIQ AHVRGHQVRK QYRAIIWSVG LLEKIILRWR RKGSGLRGFK 900 RDTITKPPEP VCPAPQEDDY DFLKEGRKQT EERLQKALTR VKSMAQYPEA RAQYRRLLTV 960 VEGFRENEAS SSSALRNNNS NTEEAANYNE EDDLIDIDSL LDDDTFMSLT FE* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
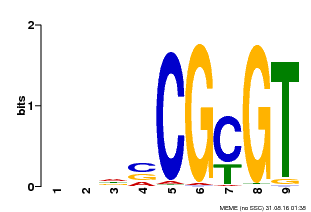
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Cagra.0248s0103.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB493811 | 0.0 | AB493811.1 Arabidopsis thaliana At5g64220 mRNA for hypothetical protein, partial cds, clone: RAAt5g64220. | |||
| GenBank | AK229403 | 0.0 | AK229403.1 Arabidopsis thaliana mRNA for Calmodulin-binding transcription activator 2, complete cds, clone: RAFL16-66-C08. | |||
| GenBank | BT010874 | 0.0 | BT010874.1 Arabidopsis thaliana At5g64220 gene, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_023637278.1 | 0.0 | calmodulin-binding transcription activator 2 isoform X2 | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | R0EUM4 | 0.0 | R0EUM4_9BRAS; Uncharacterized protein | ||||
| STRING | Cagra.0248s0103.1.p | 0.0 | (Capsella grandiflora) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4082 | 24 | 52 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Cagra.0248s0103.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



