 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cc06_g00990 | ||||||||
| Common Name | GSCOC_T00023305001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Rubiaceae; Ixoroideae; Coffeeae; Coffea
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 341aa MW: 38080.5 Da PI: 7.9571 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 101.4 | 6e-32 | 58 | 111 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
prlrWtp+LH rFv+ave+LGG+e+AtPk +l+lm++kgL ++hvkSHLQ+YR+
Cc06_g00990 58 PRLRWTPDLHLRFVHAVERLGGQERATPKLVLQLMNIKGLNIAHVKSHLQMYRS 111
8****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 3.3E-30 | 54 | 112 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 12.41 | 54 | 114 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.79E-14 | 56 | 112 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 3.3E-22 | 58 | 113 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.2E-8 | 59 | 110 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 341 aa Download sequence Send to blast |
MEGSEDTECS KACPSSGKDE VDENDGDKQK DGGSSSNSTV EENENKPSVR PYVRSKMPRL 60 RWTPDLHLRF VHAVERLGGQ ERATPKLVLQ LMNIKGLNIA HVKSHLQMYR SKKIDDPSQG 120 VTDHSHLWEG VDRNIFNLGQ IPMLPGLHQN QKSTFRYGDA SWNGLENSMQ MSAMRQSTNK 180 KRPGFYDTWP GRIVGSRNND VYSGFRNSNQ LWSWQTHELE SKMQSLPQQE YWQDQLKSSI 240 QVQLKVQDFS SCRSNIPGVG RSMIQKKRAG VKRKATDCNI HLNLSLGLGP IGDGHQDSLE 300 DDDGHLSLSL CSPSSSKITR LIDDASKENA SGASTLDLTL * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 4e-18 | 59 | 113 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 4e-18 | 59 | 113 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 4e-18 | 59 | 113 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 4e-18 | 59 | 113 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
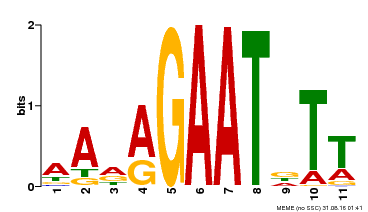
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00297 | DAP | Transfer from AT2G38300 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027066215.1 | 0.0 | uncharacterized protein LOC113692072 | ||||
| TrEMBL | A0A068UFP0 | 0.0 | A0A068UFP0_COFCA; Uncharacterized protein | ||||
| STRING | EOY07117 | 1e-100 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA4382 | 21 | 41 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G38300.1 | 2e-56 | G2-like family protein | ||||



