 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cc03_g01230 | ||||||||
| Common Name | GSCOC_T00025729001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Rubiaceae; Ixoroideae; Coffeeae; Coffea
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 676aa MW: 73886.1 Da PI: 6.5171 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 91.3 | 8.2e-29 | 217 | 270 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr++W+ eLH++Fv av+qL G++kA+Pk+ilelm+v+gLt+e+v+SHLQkYRl
Cc03_g01230 217 KPRVVWSVELHQQFVAAVNQL-GIDKAVPKKILELMNVPGLTRENVASHLQKYRL 270
79*******************.********************************8 PP
| |||||||
| 2 | Response_reg | 76.1 | 1.3e-25 | 37 | 145 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTSEEEEEESTTTHHHHHHHHHTTES CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepklpiivvtahgeeedalealkaGak 95
vl+vdD+p+ +++l+++l++ y ev+ + +e al+ll++++ +D+++ D+ mp+mdG++ll++ e +lp+i+++a ++++ + + + Ga
Cc03_g01230 37 VLVVDDDPTCLRILEKMLRNCLY-EVTKCNRAEIALKLLRDNRngYDIVISDVHMPDMDGFKLLEQVGLEM-DLPVIMMSADDSKNVVMKGVTHGAC 131
89*********************.***************999999**********************6644.8************************ PP
EEEESS--HHHHHH CS
Response_reg 96 dflsKpfdpeelvk 109
d+l Kp+ +e+l +
Cc03_g01230 132 DYLIKPVRIEALKN 145
**********9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF036392 | 1.5E-188 | 14 | 667 | IPR017053 | Response regulator B-type, plant |
| Gene3D | G3DSA:3.40.50.2300 | 7.0E-42 | 34 | 180 | No hit | No description |
| SuperFamily | SSF52172 | 7.68E-35 | 34 | 158 | IPR011006 | CheY-like superfamily |
| SMART | SM00448 | 6.8E-30 | 35 | 147 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 41.787 | 36 | 151 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 7.3E-23 | 37 | 145 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 6.80E-27 | 38 | 150 | No hit | No description |
| PROSITE profile | PS51294 | 10.896 | 214 | 273 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 5.02E-20 | 215 | 274 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.4E-30 | 216 | 275 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 9.6E-25 | 217 | 270 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 9.7E-8 | 219 | 269 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0071368 | Biological Process | cellular response to cytokinin stimulus | ||||
| GO:0080113 | Biological Process | regulation of seed growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 676 aa Download sequence Send to blast |
MNLGGGQTAK AMSAPSSSAS LKSGDVVSDQ FPAGLRVLVV DDDPTCLRIL EKMLRNCLYE 60 VTKCNRAEIA LKLLRDNRNG YDIVISDVHM PDMDGFKLLE QVGLEMDLPV IMMSADDSKN 120 VVMKGVTHGA CDYLIKPVRI EALKNIWQHV VRKRKHEWKD KDFEQSGSAE EGERLVKPLE 180 DVDYSSSATE GNWKTSKRRK DEEDEPEEKD DTSSLKKPRV VWSVELHQQF VAAVNQLGID 240 KAVPKKILEL MNVPGLTREN VASHLQKYRL YLRRLSGVSS HQNGLNSSFM GTPEATFASM 300 SSLGGLDLQA LAATGHIPAQ SLASLQALGR PTTKNSISMP MVDQRNLFSF ETPKLRFGDG 360 QQQLNNNSKQ INLLHGIPTN MEPKQLASLN QSAQTFGGLN MQVNPQASQS SSLLVQMSHP 420 MSRTQMLNEN KGNHVPRLQS SIGQPILSNG IHSGVLSRNG IVDNVRGAVY NPVSQASSLV 480 DFSINKSAEL PGNSFAQVSN SGISSLTSKG IIQEEVSSEV KGSRGFIHSY DIFNELQQHK 540 AQDWGMQNVG STFDASQHSA LQGSLDVSPS SLVQQGFSSS QKDRNVSIGD AIFSGGEGAH 600 CNNSKFGQQF NSFVADNTLR VKAERLPDSG CQNNLFPQQF GQEDLMSALL KQQEGIGSVE 660 SEFGFDGYPL DKFPV* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 3e-22 | 213 | 276 | 1 | 64 | ARR10-B |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-[AG]GATT-3'. Functions as a response regulator involved in His-to-Asp phosphorelay signal transduction system. Phosphorylation of the Asp residue in the receiver domain activates the ability of the protein to promote the transcription of target genes. Could directly activate some type-A response regulators in response to cytokinins. Involved in the expression of nuclear genes for components of mitochondrial complex I. Promotes cytokinin-mediated leaf longevity. Involved in the ethylene signaling pathway in an ETR1-dependent manner and in the cytokinin signaling pathway. {ECO:0000269|PubMed:11370868, ECO:0000269|PubMed:11574878, ECO:0000269|PubMed:15282545, ECO:0000269|PubMed:16407152}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00054 | PBM | Transfer from AT4G16110 | Download |
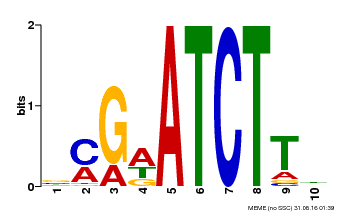
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027115945.1 | 0.0 | two-component response regulator ARR2-like | ||||
| Swissprot | Q9ZWJ9 | 0.0 | ARR2_ARATH; Two-component response regulator ARR2 | ||||
| TrEMBL | A0A068TS71 | 0.0 | A0A068TS71_COFCA; Two-component response regulator | ||||
| STRING | EMJ26354 | 0.0 | (Prunus persica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA3191 | 24 | 50 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16110.1 | 0.0 | response regulator 2 | ||||



