 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cc03_g00210 | ||||||||
| Common Name | GSCOC_T00025599001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Rubiaceae; Ixoroideae; Coffeeae; Coffea
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 985aa MW: 109622 Da PI: 7.0481 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 128.8 | 2e-40 | 133 | 210 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+C+adls+ak+yhrrhkvC+vhska+ +lv +++qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
Cc03_g00210 133 VCQVEDCRADLSNAKDYHRRHKVCDVHSKATRALVGNVMQRFCQQCSRFHVLQEFDEGKRSCRRRLAGHNRRRRKTHP 210
6**************************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 2.2E-32 | 127 | 195 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.223 | 131 | 208 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.44E-37 | 132 | 213 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 1.7E-28 | 134 | 207 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF48403 | 1.55E-9 | 708 | 864 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 3.0E-8 | 763 | 867 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.38E-8 | 763 | 864 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016021 | Cellular Component | integral component of membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 985 aa Download sequence Send to blast |
MEAKIGGKAY HYYGPVVSDL KALGKRTVEW DLNDWKWDGD LFTAAPLNSL PSDCRSRQFF 60 PTGSEIPTNS LRELEKRRRG VDGEDEELTD EAGSLHLKLG GHLYPITEGD VDKWEGKSGK 120 KTKVVGPSSN RAVCQVEDCR ADLSNAKDYH RRHKVCDVHS KATRALVGNV MQRFCQQCSR 180 FHVLQEFDEG KRSCRRRLAG HNRRRRKTHP ESVESGTPAT DERGSNYLLI SLLRILSNIH 240 SNNSSDQTKD QDLLSHLLRS LAGLAGIVNE KNLTGLLPGS QDLQNAGTSD GNPAKDPSRN 300 MLQYSTMPAS ESAQKRILGD DDNGIVRISS PAQSTLLLPP IEGILTKASS LGTTVGKTRM 360 NNIDLNNAYD DSQDCIENLQ SSDCPTHIGK ASSGCPLWVY QDPYKSSPPQ PSGNSGSTST 420 QSPSSSSGEA QSRTDRIVFK LFGKDPSDFP LALRKQILDW LSHSPSDIES YIRPGCVILT 480 IYIRMDKSTW EELCYDLTSS LRRLLDASTD SFWKSGWIYA RVRHRVAFVY DGCIVLDTPL 540 PHKSQKSCRI LNINPIAVCA SGEVKFSVRG INLSQPTTRL LCALEGKYLA QERCADVIGG 600 ADLYIEHAEI QTLNFTCTVP DVTGRGFIEV EDHGLSSSFF PFIVAENDVC SEISTLESVI 660 EAAEISNGLH GDNQNLEDRN QALDFIHEIG WLLHRSQLKF RLGQQDPNLD TFPFQRFRWL 720 IEFSVEHDWC AVVKLLLNVL FNKLMGEEKR SSIEDALLDI GLLHRAVRRN CRSMVEVLLR 780 YHPDADLNKL SPIRYVFRPD VKGPAGLTPL HIAAGRDGAE HVLDALTDDP GLVGVEAWRS 840 ARDSTGLTPN DYACLRGHYS YIHLVQKKIN KKSGSQHVVL EIPDGHLESS MNQKTADENK 900 ARKVSSLSTE MSVVKPSHVH CRQCEQKLAY GRNRTSLAIY RPAMLSMVAI AAVCVCVALL 960 FKSSPEVDYV YGPFRWEYLE YGSS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 5e-30 | 128 | 207 | 5 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16095614}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
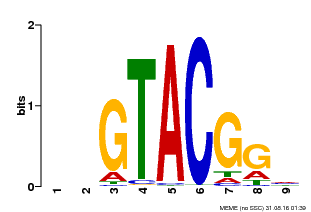
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027115807.1 | 0.0 | LOW QUALITY PROTEIN: squamosa promoter-binding-like protein 1 | ||||
| Swissprot | Q9SMX9 | 0.0 | SPL1_ARATH; Squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | A0A068TRT4 | 0.0 | A0A068TRT4_COFCA; Uncharacterized protein | ||||
| STRING | POPTR_0008s09810.1 | 0.0 | (Populus trichocarpa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA1871 | 24 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



