 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cc01_g07440 | ||||||||
| Common Name | GSCOC_T00030182001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Rubiaceae; Ixoroideae; Coffeeae; Coffea
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 523aa MW: 56969.5 Da PI: 5.9703 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 56.1 | 8.7e-18 | 39 | 86 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT Ed +lv++v+++G g+W+++ ++ g+ R++k+c++rw ++l
Cc01_g07440 39 KGPWTSAEDAILVEYVTKHGEGNWNAVQKHSGLARCGKSCRLRWANHL 86
79******************************************9986 PP
| |||||||
| 2 | Myb_DNA-binding | 52.3 | 1.3e-16 | 92 | 135 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
+g++T+eE+ +++++++++G++ W++ a++++ gRt++++k++w++
Cc01_g07440 92 KGAFTPEEERRIIELHAKMGNK-WARMAAELP-GRTDNEIKNYWNT 135
799*******************.*********.***********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 16.581 | 34 | 86 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 3.26E-30 | 37 | 133 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.7E-13 | 38 | 88 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.9E-15 | 39 | 86 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.3E-24 | 40 | 93 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 4.87E-11 | 41 | 86 | No hit | No description |
| PROSITE profile | PS51294 | 26.474 | 87 | 141 | IPR017930 | Myb domain |
| SMART | SM00717 | 1.8E-16 | 91 | 139 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.5E-15 | 92 | 135 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 5.1E-26 | 94 | 140 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 3.78E-12 | 94 | 135 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009789 | Biological Process | positive regulation of abscisic acid-activated signaling pathway | ||||
| GO:0043068 | Biological Process | positive regulation of programmed cell death | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0045926 | Biological Process | negative regulation of growth | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 523 aa Download sequence Send to blast |
MSMTSESEDR MVPKGAIGSP SIEETSGGGN FGVNGPLKKG PWTSAEDAIL VEYVTKHGEG 60 NWNAVQKHSG LARCGKSCRL RWANHLRPDL KKGAFTPEEE RRIIELHAKM GNKWARMAAE 120 LPGRTDNEIK NYWNTRIKRR QRAGLPIYPP DICLQALSES KQSENLSSFS SVETQHPDLL 180 PINSFEIPAV EFKNLELNHQ LYPSPLLDVP GRGLLDIPAS SLLAQGLHSS YGGKTLLSAV 240 HPTKRLRQSE SLFPGLATSF TGAFTNCSKY HNDGSAQSLQ SFGITSAYDQ NLTSDNHSTS 300 CVLPGSHALL NGNPSSSEPS WAMKLELPSL QTQIGSWGSP SSPLPSLESV DTLIQSPPTE 360 HTQSGSLSPR NSGLLDAVLY ESQSLKNFKS SSCQHMSSVS MAQGEVMDNS LQDLHEAEWE 420 AYGDPISPRG HSAASVLSEY TPISRSSMDE LQSLENTPGG KVKQEAEEMV PTQFCGNDDA 480 FNSNNMIFSR PDFLLASNCF GPKGEYGKEQ LHAERCSWGT SW* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 1e-31 | 37 | 140 | 5 | 107 | B-MYB |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator of gibberellin-dependent alpha-amylase expression in aleurone cells. Involved in pollen and floral organs development. May bind to the 5'-TAACAAA-3' box of alpha-amylase promoter. {ECO:0000269|PubMed:9150608}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00490 | DAP | Transfer from AT5G06100 | Download |
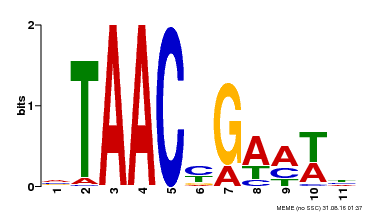
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By gibberellin in aleurone cells. {ECO:0000269|PubMed:9150608}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027081255.1 | 0.0 | transcription factor GAMYB-like | ||||
| Swissprot | A2WW87 | 1e-102 | GAM1_ORYSI; Transcription factor GAMYB | ||||
| TrEMBL | A0A068UQ74 | 0.0 | A0A068UQ74_COFCA; Uncharacterized protein | ||||
| STRING | XP_009786742.1 | 0.0 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA2021 | 23 | 58 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06100.1 | 9e-88 | myb domain protein 33 | ||||



