 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Ciclev10026023m | ||||||||
| Common Name | CICLE_v10026023mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Rutaceae; Aurantioideae; Citrus
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 336aa MW: 37120.8 Da PI: 7.0925 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 58.7 | 1.3e-18 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT +Ed++lv++++++G g+W++ ++ g++R++k+c++rw +yl
Ciclev10026023m 14 KGPWTSDEDQKLVKYIQKHGHGSWRALPKLAGLNRCGKSCRLRWTNYL 61
79********************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 54.9 | 2.1e-17 | 67 | 112 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg++++eE++ +++++ lG++ W++Ia +++ gRt++++k++w+++l
Ciclev10026023m 67 RGKFSQEEEQTILNLHSILGNK-WSAIAGHLP-GRTDNEIKNFWNTHL 112
89********************.*********.************996 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.3E-25 | 5 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 17.687 | 9 | 61 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 6.6E-32 | 11 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 6.3E-15 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.2E-17 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.60E-11 | 16 | 61 | No hit | No description |
| PROSITE profile | PS51294 | 26.442 | 62 | 116 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.1E-27 | 65 | 117 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.7E-16 | 66 | 114 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 6.0E-16 | 67 | 112 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 8.32E-12 | 69 | 112 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 336 aa Download sequence Send to blast |
MGRSPCCDES GLKKGPWTSD EDQKLVKYIQ KHGHGSWRAL PKLAGLNRCG KSCRLRWTNY 60 LRPDIKRGKF SQEEEQTILN LHSILGNKWS AIAGHLPGRT DNEIKNFWNT HLKKKLIQMG 120 FDPMTHRPRT DVFSSLPHLI ALANLKELME NHTWEEQAVR LQAEMTKLQY LQYLLQPSSS 180 NSLNNSSSSA FTDIDSINLF NSLSSSTQWD MGSASLQNLS NPSVSFSHMP DLQVPCSYQT 240 PLNKNSTDNT VHGAEFPLLS QGESFSPKSP WQQLAAASSA TPSPPSMAPP PLTESSINNL 300 VDACSTSSSY GGSVVAAPSV WPELLLEDPL FHEIA* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 5e-30 | 12 | 116 | 5 | 108 | B-MYB |
| Search in ModeBase | ||||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Highly expressed in roots and at lower levels in stems, flowers and siliques. {ECO:0000269|PubMed:24902892}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250|UniProtKB:Q9M2D9}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00503 | DAP | Transfer from AT5G10280 | Download |
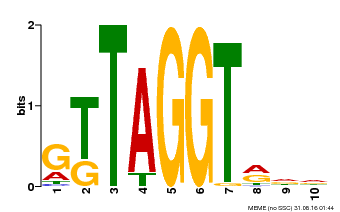
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by gravity in roots. {ECO:0000269|PubMed:24902892}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AM430933 | 7e-41 | AM430933.2 Vitis vinifera contig VV78X227166.3, whole genome shotgun sequence. | |||
| GenBank | AM442163 | 7e-41 | AM442163.1 Vitis vinifera, whole genome shotgun sequence, contig VV78X189718.9, clone ENTAV 115. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006428207.1 | 0.0 | transcription factor MYB93 | ||||
| Refseq | XP_006464255.1 | 0.0 | transcription factor MYB93 | ||||
| Swissprot | Q9SBF3 | 1e-106 | MYB92_ARATH; Transcription factor MYB92 | ||||
| TrEMBL | V4SLA6 | 0.0 | V4SLA6_9ROSI; Uncharacterized protein | ||||
| STRING | XP_006464255.1 | 0.0 | (Citrus sinensis) | ||||
| STRING | XP_006428207.1 | 0.0 | (Citrus clementina) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4 | 28 | 2646 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G34670.1 | 7e-97 | myb domain protein 93 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Ciclev10026023m |
| Entrez Gene | 18037080 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



