 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Ciclev10018744m | ||||||||
| Common Name | CICLE_v10018744mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Rutaceae; Aurantioideae; Citrus
|
||||||||
| Family | Nin-like | ||||||||
| Protein Properties | Length: 944aa MW: 104105 Da PI: 5.6277 | ||||||||
| Description | Nin-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | RWP-RK | 92.1 | 4.2e-29 | 536 | 586 | 2 | 52 |
RWP-RK 2 ekeisledlskyFslpikdAAkeLgvclTvLKriCRqyGIkRWPhRkiksl 52
ek+isle+l++yF++++kdAAk+Lgvc+T++KriCRq+GI+RWP+Rki+++
Ciclev10018744m 536 EKSISLEVLQQYFAGSLKDAAKSLGVCPTTMKRICRQHGISRWPSRKINKV 586
799**********************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51519 | 17.575 | 525 | 606 | IPR003035 | RWP-RK domain |
| Pfam | PF02042 | 3.6E-26 | 539 | 586 | IPR003035 | RWP-RK domain |
| SuperFamily | SSF54277 | 3.92E-23 | 842 | 931 | No hit | No description |
| SMART | SM00666 | 9.1E-26 | 847 | 929 | IPR000270 | PB1 domain |
| PROSITE profile | PS51745 | 25.265 | 847 | 929 | IPR000270 | PB1 domain |
| Gene3D | G3DSA:3.10.20.240 | 2.2E-27 | 847 | 928 | No hit | No description |
| CDD | cd06407 | 5.53E-39 | 848 | 928 | No hit | No description |
| Pfam | PF00564 | 8.2E-19 | 848 | 928 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0010167 | Biological Process | response to nitrate | ||||
| GO:0042128 | Biological Process | nitrate assimilation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 944 aa Download sequence Send to blast |
MSPFLISEQP CSPLWAFSDA DNDDKLSGHV NYPLFLKCNP NSETENPKDN DENRRFPSPL 60 SALMPLENPD GYCMIKERIT QALRYFKDST EQHVLAQVWV PVKVGGRYVL TTSGQPFVLD 120 PHSNGLHQYR MVSLMYMFSV DGESDGELGL PGRVFWQKLP EWTPNVQYYS SKEYSRLDHA 180 LHHNVRGTMA LPVFEPSGQS CVAVIELIMT SQKINYAPEV DKVCKALEAV NLKSSEILDY 240 PSTQICNEGR QNALAEILEI LSVVCETHKL PLAQTWVPCR HRSVLAYGGG LKKSCSSIDG 300 SCMGQVCMST TDVAFYVVDG HMWGFREACV EHHLQKDQGV AGRAFFSLSS CFCKDITQFC 360 KTEYPLVHYA RMFGLTSCFA ICLRSTYTGD DDYILEFFLP PAITDSYEQQ TLLGSILATM 420 KQHFQSLKVA SGIDLEDDEG TIEIIEGEAT ADKKLNLRME SIRIPQSVRS PPQPHALPNG 480 GELGQLDIPE QQLMENFDYM NSRGNAVNVG GNDNPVSLLE NKNTRKPSER KRGKTEKSIS 540 LEVLQQYFAG SLKDAAKSLG VCPTTMKRIC RQHGISRWPS RKINKVNRSL TKLKRVIESV 600 QGTNGTFGLT SLTTSPLPVA VNSISWPSGL NGSNQQNSPN SKPELLGEKI LSPIYKTPGS 660 DGHTELEDRL SGGRMSTHEE HIHEQNALSP EIGKGKNSPK TGSGSREESA GSPTSHGSCQ 720 GNPANESAPA KDVLVSSIHE PRFKVGGSLE LVFQPVGEMN LSAAFSIPDA LVTTEPQEPF 780 GGLLVEDAGS SKDLRNLCPA VADAIVDERL PENSCANLPC AELSPKQHLA TLSQTMPRVY 840 SRQEMKSVTI KATYREDIIR FRISLSCGIL ELKEEVAKRL KLELGTFDIK YLDDDQEWVL 900 IACDADLQEC LDISRSSGSN MIRLSIHDIM ANLGSSCEST GEL* |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, leaves, flowers and siliques. Detected in root hairs, emerging secondary roots, vascular tissues, leaf parenchyma cells and stomata. {ECO:0000269|PubMed:18826430}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in regulation of nitrate assimilation and in transduction of the nitrate signal. {ECO:0000269|PubMed:18826430}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00449 | DAP | Transfer from AT4G24020 | Download |
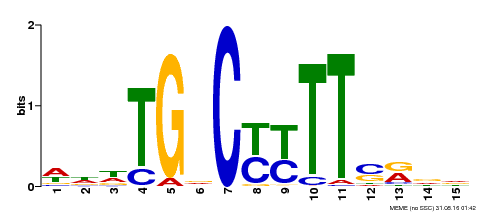
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Not regulated by the N source or by nitrate. {ECO:0000269|PubMed:18826430}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KX056270 | 0.0 | KX056270.1 Citrus trifoliata transcription factor NLP7 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006439290.2 | 0.0 | LOW QUALITY PROTEIN: protein NLP7 | ||||
| Swissprot | Q84TH9 | 0.0 | NLP7_ARATH; Protein NLP7 | ||||
| TrEMBL | V4TLI6 | 0.0 | V4TLI6_9ROSI; Uncharacterized protein | ||||
| STRING | XP_006439290.1 | 0.0 | (Citrus clementina) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2507 | 26 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G24020.1 | 0.0 | NIN like protein 7 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Ciclev10018744m |
| Entrez Gene | 18047473 |



