 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Ciclev10018697m | ||||||||
| Common Name | CICLE_v10018697mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Rutaceae; Aurantioideae; Citrus
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 989aa MW: 109658 Da PI: 6.482 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 132.4 | 1.6e-41 | 149 | 226 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+C adls+ak+yhrrhkvCe+hska+ +lv +++qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+++
Ciclev10018697m 149 VCQVEDCGADLSNAKDYHRRHKVCEMHSKASRALVGNVMQRFCQQCSRFHVLQEFDEGKRSCRRRLAGHNKRRRKTNP 226
6**************************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 9.2E-34 | 142 | 211 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.385 | 147 | 224 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 6.67E-39 | 148 | 229 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 8.6E-30 | 150 | 223 | IPR004333 | Transcription factor, SBP-box |
| Gene3D | G3DSA:1.25.40.20 | 5.3E-5 | 304 | 333 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 8.24E-6 | 761 | 873 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 5.3E-5 | 762 | 872 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 2.69E-6 | 762 | 873 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016021 | Cellular Component | integral component of membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 989 aa Download sequence Send to blast |
METRFRGEAH HFYGMNSMDL RAVGKKTLEW DLNDWKWDGD LFIASKLNPA PNENIGRQFF 60 PLAVGNSSNS SSSCSDEVNL GIENGKREVE KKRRAVVVED HNSYEVAAGG LSLKLGGNGH 120 PLSEREMGNW AGSSGKKTKF GGGSSSRAVC QVEDCGADLS NAKDYHRRHK VCEMHSKASR 180 ALVGNVMQRF CQQCSRFHVL QEFDEGKRSC RRRLAGHNKR RRKTNPDAVA NGSSPNNDQT 240 SGYLLISLLR ILSNMHSSRS DQRTDQDLLS HLLRGLASPA GENGGRGISG LLQEHQDMLN 300 ERTSAGNSEV VQAFLANGQG CPTPFRQQLN ATVSEMPQQV SLPHDARGAE DQDGNVAQIK 360 MNNFDLNDVY IDSDDGTEDV ERSPVPANLG TSSIDCPSWV RQDSQQSSPP QTSGNSDSAS 420 AQSPSSSSDA QSRTDRIVFK LFGKEPNDFP LVLRAQILDW LSHSPSDMES YIRPGCVILT 480 IYLRQAEAAW EELCCDLTFS LSRLLDLSND SFWTSGWVYA RVQHQIAFIY NGQVVLDTSL 540 PPRSNNYSKI LSVKPIAVPA SERAQFFVKG INLGRSATRL LCAVEGKYMV QEATHELLDD 600 VDGFKELDEL QCVNFSCSIP AVTGRGFIEI EDHGFSSTFF PFIVAEEDVC SEIRMLESAL 660 EFNRTDADVE RFGKIDTKNQ AMDFIHEIGW LFHRSQSKSR LGHLDPNTDL FPLRRFKWLI 720 EFSMDHEWCA VVKKLLHILL DGTVSLGEHP SLDLALTELG LLHRAVRKNS RPLVDLLLRF 780 VPLEVSDRLG SENKALVDGV HKGFLFRPDV IGPAGLTPIH IAAGKDGSED VLDALTDDPG 840 MVGIEAWKNA RDSSGSTPED YARLRGHYSY IHLVQKKINK RPNGGHVVVD ICGVVPDSNI 900 YQKQNNESTA SFEIGQTPVR PTQHNCKLCH QKLGYATASR SLVYKPAMLS MVAIAAVCVC 960 VALLFKSCPE VLYVFRPFRW EMLDYGTS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 2e-31 | 143 | 223 | 4 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Expressed during plant development. {ECO:0000269|PubMed:10524240}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000269|PubMed:16554053}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
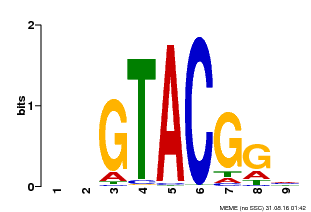
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006444595.1 | 0.0 | squamosa promoter-binding-like protein 1 | ||||
| Swissprot | Q9S7P5 | 0.0 | SPL12_ARATH; Squamosa promoter-binding-like protein 12 | ||||
| TrEMBL | A0A067H4P9 | 0.0 | A0A067H4P9_CITSI; Uncharacterized protein | ||||
| TrEMBL | V4U1C3 | 0.0 | V4U1C3_9ROSI; Squamosa promoter-binding protein 5 | ||||
| STRING | XP_006444595.1 | 0.0 | (Citrus clementina) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3030 | 27 | 64 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Ciclev10018697m |
| Entrez Gene | 18048503 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



