 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Ciclev10012152m | ||||||||
| Common Name | CICLE_v10012152mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Rutaceae; Aurantioideae; Citrus
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 337aa MW: 37078.3 Da PI: 6.3685 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 55 | 1.8e-17 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+eEd +lv +++++G+g+W++++ g+ R+ k+c++rw +yl
Ciclev10012152m 14 KGPWTPEEDIILVSYIQEHGPGNWRAVPTNTGLLRCSKSCRLRWTNYL 61
79******************************99************97 PP
| |||||||
| 2 | Myb_DNA-binding | 49.2 | 1.3e-15 | 67 | 112 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg++T +E++ ++++ ++lG++ W++Ia++++ Rt++++k++w+++l
Ciclev10012152m 67 RGNFTDQEEKMIIHLQALLGNR-WAAIASYLP-QRTDNDIKNYWNTHL 112
89********************.*********.************996 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 24.604 | 9 | 65 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.4E-30 | 11 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.0E-14 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.4E-16 | 14 | 61 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.3E-23 | 15 | 68 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.54E-11 | 16 | 61 | No hit | No description |
| PROSITE profile | PS51294 | 20.048 | 66 | 116 | IPR017930 | Myb domain |
| SMART | SM00717 | 1.8E-15 | 66 | 114 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.2E-14 | 67 | 112 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.38E-11 | 69 | 112 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 1.4E-24 | 69 | 116 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0001666 | Biological Process | response to hypoxia | ||||
| GO:0009617 | Biological Process | response to bacterium | ||||
| GO:0009626 | Biological Process | plant-type hypersensitive response | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0042761 | Biological Process | very long-chain fatty acid biosynthetic process | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 337 aa Download sequence Send to blast |
MGRPPCCDKI GIKKGPWTPE EDIILVSYIQ EHGPGNWRAV PTNTGLLRCS KSCRLRWTNY 60 LRPGIKRGNF TDQEEKMIIH LQALLGNRWA AIASYLPQRT DNDIKNYWNT HLKKKVRKLQ 120 LAAAGCSEDN SQYRDELASA SSQQISRGQW ERRLQTDIHM AKQALCAALS PDKASILSEL 180 KPANGFISYT KPAVQAPAYA SSTENIAKLL KGWTRNAQKS ASSNSGVTDQ NSINNNVNHI 240 AGAESASSEE TPSKVASNST GIELSEAFES LFGFESFDSS NSTDLSQSVT PESSTFQDYE 300 SKQLLLDPSA GADDDQMPQL SLLEKWLFDD QGGKDI* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1h8a_C | 7e-26 | 12 | 116 | 25 | 128 | MYB TRANSFORMING PROTEIN |
| Search in ModeBase | ||||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Expression in flowers increases as the flowers develop. {ECO:0000269|PubMed:1840903}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in flowers, leaves and weakly in seed pods. {ECO:0000269|PubMed:1840903}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. {ECO:0000305}. | |||||
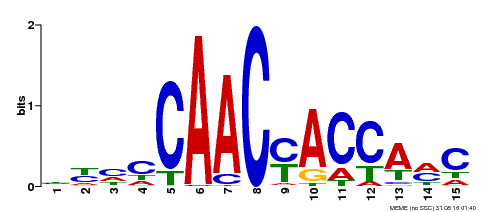
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00386 | DAP | Transfer from AT3G28910 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | EF071983 | 0.0 | EF071983.1 Citrus macrophylla MYB60-like protein gene, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006431144.1 | 0.0 | myb-related protein 306 | ||||
| Swissprot | P81392 | 1e-116 | MYB06_ANTMA; Myb-related protein 306 | ||||
| TrEMBL | V4UX79 | 0.0 | V4UX79_9ROSI; Uncharacterized protein | ||||
| STRING | XP_006431144.1 | 0.0 | (Citrus clementina) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4 | 28 | 2646 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G28910.1 | 1e-112 | myb domain protein 30 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Ciclev10012152m |
| Entrez Gene | 18039235 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



