 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Ciclev10000756m | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Rutaceae; Aurantioideae; Citrus
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 552aa MW: 60220 Da PI: 4.9121 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 52.9 | 8.5e-17 | 38 | 85 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT Ed +l+d+vk++G g+W+++ + g+ R++k+c++rw ++l
Ciclev10000756m 38 KGPWTSAEDAILIDYVKKHGEGNWNAVQKNSGLFRCGKSCRLRWANHL 85
79******************************************9986 PP
| |||||||
| 2 | Myb_DNA-binding | 52.7 | 1e-16 | 91 | 134 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
+g++T+eE++ +v++++++G++ W++ a++++ gRt++++k++w++
Ciclev10000756m 91 KGAFTQEEEQMIVELHAKMGNK-WARMAAHLP-GRTDNEIKNYWNT 134
799*******************.*********.***********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 15.816 | 33 | 85 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 4.87E-30 | 37 | 132 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 8.9E-15 | 37 | 87 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.1E-14 | 38 | 85 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.4E-23 | 39 | 92 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.90E-11 | 40 | 85 | No hit | No description |
| PROSITE profile | PS51294 | 26.169 | 86 | 140 | IPR017930 | Myb domain |
| SMART | SM00717 | 7.8E-17 | 90 | 138 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.2E-15 | 91 | 134 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.70E-12 | 93 | 134 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 4.1E-26 | 93 | 139 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009789 | Biological Process | positive regulation of abscisic acid-activated signaling pathway | ||||
| GO:0043068 | Biological Process | positive regulation of programmed cell death | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0045926 | Biological Process | negative regulation of growth | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 552 aa Download sequence Send to blast |
MSCSSNGSDD MMLSNDQTDS PLMDEGGGGT ARGVILKKGP WTSAEDAILI DYVKKHGEGN 60 WNAVQKNSGL FRCGKSCRLR WANHLRPNLK KGAFTQEEEQ MIVELHAKMG NKWARMAAHL 120 PGRTDNEIKN YWNTRIKRRQ RAGLPLYPPE VSFQALQESQ CQNINGINSG DKGHHDLLQA 180 NGYEIPDVVF DSLKANQGVL PYVPDIPDIT ASTMLMKGLG SQYCSFVPPV MHRPKRLRES 240 VPLFSGYGGS VKNEFSFDQF HNDTSDKIAR SFGLSFPLDP DTTPKAPESF GMLQGSHTLS 300 NGNFSASKPN LGAVKLELPS LQYTETDLGS WGTSSPQPLL ETVDAFIQSP PPTGTLDSGC 360 PSPRNSGLLD ALIHESNILK NHSSDKSSNS SSVTPDVADS CTRNFCETEW EDYSDPLSPL 420 GHSATSLFNE CTPLSASGSS LDEQAPVETF GYKVKSEPVD QAWTPDREKE SSNLYVSQPD 480 ALLASDWFEH GSSYVKDPIV MNDAIATLLG DDLRSEYKQI STGTSAPSQG WGFSSCAWNN 540 MPAVCQMSEL P* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 6e-31 | 36 | 139 | 5 | 107 | B-MYB |
| Search in ModeBase | ||||||
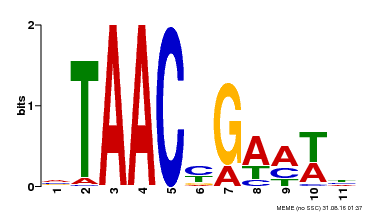
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00490 | DAP | Transfer from AT5G06100 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006432323.2 | 0.0 | transcription factor MYB33 | ||||
| Refseq | XP_006471261.1 | 0.0 | transcription factor MYB33 | ||||
| Refseq | XP_015383648.1 | 0.0 | transcription factor MYB33 | ||||
| Refseq | XP_024040037.1 | 0.0 | transcription factor MYB33 | ||||
| Refseq | XP_024040038.1 | 0.0 | transcription factor MYB33 | ||||
| Refseq | XP_024951612.1 | 0.0 | transcription factor MYB33 | ||||
| Refseq | XP_024951613.1 | 0.0 | transcription factor MYB33 | ||||
| Refseq | XP_024951614.1 | 0.0 | transcription factor MYB33 | ||||
| TrEMBL | A0A2H5PPC7 | 0.0 | A0A2H5PPC7_CITUN; Uncharacterized protein | ||||
| STRING | XP_006471261.1 | 0.0 | (Citrus sinensis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3802 | 28 | 59 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06100.3 | 1e-143 | myb domain protein 33 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Ciclev10000756m |



