 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Ciclev10000491m | ||||||||
| Common Name | CICLE_v10000491mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Rutaceae; Aurantioideae; Citrus
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 683aa MW: 75901.4 Da PI: 6.2389 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 71.2 | 1.3e-22 | 131 | 232 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SS CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgr 87
f+k+lt sd++++g +++ +++a+++ +++ ++l+ +d++g++W +++i+r++++r++lt+GW+ Fv++++L +gD ++F +
Ciclev10000491m 131 FCKTLTASDTSTHGGFSVLRRHADDClppldmsQQP--PWQELVATDLHGNEWHFRHIFRGQPRRHLLTTGWSVFVSSKKLVAGDAFIFL--RG 220
99****************************964333..4459************************************************..34 PP
SEE..EEEEE-S CS
B3 88 sefelvvkvfrk 99
+++el+v+v+r+
Ciclev10000491m 221 ENEELRVGVRRH 232
88889****996 PP
| |||||||
| 2 | Auxin_resp | 111.8 | 6.8e-37 | 257 | 339 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
a+ha++t++ F+++Y+Pr+s+seF+v+v+k+++a ++k+svGmRfkm fe+e+++err+sGt+vgv d+ ++ W++S+WrsLk
Ciclev10000491m 257 ASHAIATGTLFSIFYKPRTSRSEFIVSVNKYLEAKRHKLSVGMRFKMTFEGEEVPERRFSGTIVGVGDCRSSVWQDSEWRSLK 339
789******************************************************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 3.79E-44 | 121 | 260 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 3.6E-41 | 128 | 246 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 2.42E-20 | 129 | 231 | No hit | No description |
| SMART | SM01019 | 3.1E-22 | 131 | 233 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 1.9E-20 | 131 | 232 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 11.124 | 131 | 233 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 1.1E-34 | 257 | 339 | IPR010525 | Auxin response factor |
| Pfam | PF02309 | 2.4E-12 | 555 | 648 | IPR033389 | AUX/IAA domain |
| PROSITE profile | PS51745 | 28.221 | 558 | 651 | IPR000270 | PB1 domain |
| SuperFamily | SSF54277 | 6.15E-12 | 560 | 635 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0042802 | Molecular Function | identical protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 683 aa Download sequence Send to blast |
MAFAAAAAPL NHASGGPRTG GSNDALYREL WHACAGPLVT LPREGERVYY FPQGHMEQLE 60 ASMHQGLEQQ MPSFNLPSKI LCKVVNVQRR AEPETDEVYA QITLLPEPDQ SEVTSPDPPI 120 PEPEKCTVHS FCKTLTASDT STHGGFSVLR RHADDCLPPL DMSQQPPWQE LVATDLHGNE 180 WHFRHIFRGQ PRRHLLTTGW SVFVSSKKLV AGDAFIFLRG ENEELRVGVR RHMRQQTNMP 240 SSVISSHSMH LGVLATASHA IATGTLFSIF YKPRTSRSEF IVSVNKYLEA KRHKLSVGMR 300 FKMTFEGEEV PERRFSGTIV GVGDCRSSVW QDSEWRSLKV QWDEPSSILR PERVSSWELE 360 PLVATTTTTN AQPAQRNKRS RPPVLPSPAA DISVLGMWKS HVESSAFSYC DSPQGRDLYP 420 SPKFSTATKG NHFGLSGNKS LAAVSSNSMY WPNRAENVTE SFAPVVNKEC SEKRQGNGNT 480 CRLFGIQLVD NSNVEEASPA FTMSGAMGDD RAIPCLDADS DQHSEPSNIN RSDVPSVSCD 540 AEKSCLRSPQ ESQSRQIRSC TKVHMQGIAV GRAVDLTRFD CYEDLLKKLE EMFDIEGDLC 600 GSTKKWQVVY TDDEDDMMMV GDDPWHEFCS MVRKIFIYTA EEVKKLSPKV KLPGNEELNA 660 GKPDVEVVVN TDDRSSIVGP GC* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 0.0 | 8 | 362 | 2 | 355 | Auxin response factor 1 |
| 4ldw_A | 0.0 | 8 | 362 | 2 | 355 | Auxin response factor 1 |
| 4ldw_B | 0.0 | 8 | 362 | 2 | 355 | Auxin response factor 1 |
| 4ldx_A | 0.0 | 8 | 362 | 2 | 355 | Auxin response factor 1 |
| 4ldx_B | 0.0 | 8 | 362 | 2 | 355 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Expressed in the sepals and carpels of young flower buds. At stage 10 of flower development, expression in the carpels becomes restricted to the style. Also expressed in anthers and filaments. At stage 13, expressed in the region at the top of the pedicel, including the abscission zone. {ECO:0000269|PubMed:16176952}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in the whole plant. {ECO:0000269|PubMed:10476078}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). Seems to act as transcriptional repressor. Formation of heterodimers with Aux/IAA proteins may alter their ability to modulate early auxin response genes expression. Promotes flowering, stamen development, floral organ abscission and fruit dehiscence. Acts as repressor of IAA2, IAA3 and IAA7. {ECO:0000269|PubMed:12036261, ECO:0000269|PubMed:16176952}. | |||||
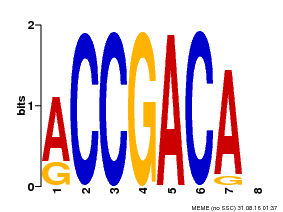
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00097 | PBM | Transfer from AT1G59750 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006431443.1 | 0.0 | auxin response factor 1 isoform X1 | ||||
| Refseq | XP_024039738.1 | 0.0 | auxin response factor 1 isoform X1 | ||||
| Refseq | XP_024039739.1 | 0.0 | auxin response factor 1 isoform X1 | ||||
| Swissprot | Q8L7G0 | 0.0 | ARFA_ARATH; Auxin response factor 1 | ||||
| TrEMBL | V4SAY2 | 0.0 | V4SAY2_9ROSI; Auxin response factor | ||||
| STRING | XP_006431443.1 | 0.0 | (Citrus clementina) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM8825 | 22 | 27 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G59750.4 | 0.0 | auxin response factor 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Ciclev10000491m |
| Entrez Gene | 18040560 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



