 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | C.cajan_40853 | ||||||||
| Common Name | KK1_041434 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Cajanus
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 388aa MW: 43966.1 Da PI: 8.5488 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 102.2 | 3.4e-32 | 66 | 119 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
prlrWtp+LH rFv+av++LGG+e+AtPk +l+lm++kgL+++hvkSHLQ+YR+
C.cajan_40853 66 PRLRWTPDLHLRFVHAVQRLGGQERATPKLVLQLMNIKGLSIAHVKSHLQMYRS 119
8****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 2.2E-29 | 62 | 120 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 13.182 | 62 | 122 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.42E-14 | 65 | 120 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 6.2E-23 | 66 | 121 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.5E-9 | 67 | 118 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 388 aa Download sequence Send to blast |
MEEEGHGSEC YKTSPSNEED YCEESEANNE DRSKQKNNGG SSSNSTVEEN EKKTTTVRPY 60 VRSKLPRLRW TPDLHLRFVH AVQRLGGQER ATPKLVLQLM NIKGLSIAHV KSHLQMYRSK 120 KIDTNQVLGD PRLLVETGDK NVYKLSQLPM LQGYNPKQSS AYRQLNFNFK FFITCIKNCT 180 TEYGYGDASL AIYENMVHRP FLSRSTSVDE SGPEFYGSIL SERIHGTNSN INWTHNKFQV 240 DSSRFNEMST NKVHEQKDKF FSFGGHESSC TQIKMSQVGL NQSTQQPSAQ QLMPYNKLAT 300 NPVELKTLKR KASDIDHLDL NLSLKINSRV VSDENQGSKS MVDHEVDSNL SLSLCSQSSF 360 SHLSNRLRRA QDHSKGQGER ASALDLTI |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 1e-17 | 67 | 121 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 1e-17 | 67 | 121 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 1e-17 | 67 | 121 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 1e-17 | 67 | 121 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 1e-17 | 67 | 121 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 1e-17 | 67 | 121 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 1e-17 | 67 | 121 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 1e-17 | 67 | 121 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 1e-17 | 67 | 121 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 1e-17 | 67 | 121 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 1e-17 | 67 | 121 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 1e-17 | 67 | 121 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00297 | DAP | Transfer from AT2G38300 | Download |
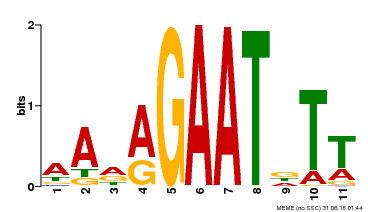
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | C.cajan_40853 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015037 | 2e-72 | AP015037.1 Vigna angularis var. angularis DNA, chromosome 4, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020204876.1 | 0.0 | two-component response regulator ARR14 | ||||
| TrEMBL | A0A151R444 | 0.0 | A0A151R444_CAJCA; Putative Myb family transcription factor At1g14600 family | ||||
| STRING | GLYMA18G43550.2 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2378 | 34 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G38300.1 | 4e-49 | G2-like family protein | ||||



