 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | C.cajan_27183 | ||||||||
| Common Name | KK1_030228 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Cajanus
|
||||||||
| Family | WRKY | ||||||||
| Protein Properties | Length: 510aa MW: 56077.8 Da PI: 6.4174 | ||||||||
| Description | WRKY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | WRKY | 83.6 | 1.9e-26 | 104 | 160 | 2 | 59 |
--SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 2 dDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhe 59
+DgynWrKYGqK+vkg+ef rsYY+Ct+++C++kk++++s+ ++++++ + g+Hnh+
C.cajan_27183 104 KDGYNWRKYGQKHVKGNEFIRSYYKCTHPNCQAKKQLQQSN-NGHITDSICIGQHNHP 160
7***************************************9.***************8 PP
| |||||||
| 2 | WRKY | 104.2 | 7.2e-33 | 289 | 347 | 1 | 59 |
---SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 1 ldDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhe 59
++Dgy+WrKYGqK vkg+++prsYYrC+ +gCpvkk+ver+++d k+v++tYeg+H+he
C.cajan_27183 289 VNDGYRWRKYGQKLVKGNTNPRSYYRCSNPGCPVKKHVERASHDSKIVITTYEGQHDHE 347
58********************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.20.25.80 | 2.0E-23 | 97 | 162 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 19.187 | 98 | 162 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 2.35E-22 | 99 | 162 | IPR003657 | WRKY domain |
| SMART | SM00774 | 9.0E-29 | 103 | 161 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 2.7E-21 | 104 | 160 | IPR003657 | WRKY domain |
| Gene3D | G3DSA:2.20.25.80 | 7.0E-35 | 275 | 349 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 1.11E-28 | 281 | 349 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 34.612 | 284 | 349 | IPR003657 | WRKY domain |
| SMART | SM00774 | 1.8E-37 | 289 | 348 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 8.2E-26 | 290 | 346 | IPR003657 | WRKY domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009863 | Biological Process | salicylic acid mediated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 510 aa Download sequence Send to blast |
MVSSDKSADQ NIPSDRLQQR VSPDSDITLS QGHDTKNELS KPEGATSIPS IVAKNEGKDS 60 DQEGSTCSLP LEKHLQSPDT LSHELPPLQS NQESPSIIRE KVSKDGYNWR KYGQKHVKGN 120 EFIRSYYKCT HPNCQAKKQL QQSNNGHITD SICIGQHNHP RPQLNSTVSV ECVLPIVEQT 180 PPKPSLADVE DPNVFYDIIH EDKTSIEQGC MPQQIKPLQS IPTTKVSPVI ELKAAHLQLT 240 KAKNQAHGNE DPESKRLKKN NSNADATGVD MSTCESRVVV QTSSEVDLVN DGYRWRKYGQ 300 KLVKGNTNPR SYYRCSNPGC PVKKHVERAS HDSKIVITTY EGQHDHEIPP ARTVTHNAAT 360 NTHTTTTNGE AGTKSGGNTV FVDTGECSDL DSVSRLTEQL NDKSNTKSKD GDKLEFRVIS 420 LSNEGPEIKF SKQQQQKDNS GAKNNSVSND IVCHSSSGVP CRSNEQLEDE VKTISEGSKD 480 CLNVVAVCDT PSTENEFNKQ SAADAEPVQS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2ayd_A | 1e-36 | 276 | 351 | 1 | 76 | WRKY transcription factor 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00094 | SELEX | Transfer from AT2G04880 | Download |
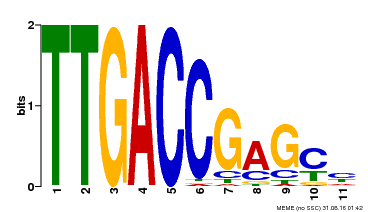
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | C.cajan_27183 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015036 | 1e-139 | AP015036.1 Vigna angularis var. angularis DNA, chromosome 3, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020234360.1 | 0.0 | WRKY transcription factor 1 | ||||
| TrEMBL | A0A151S002 | 0.0 | A0A151S002_CAJCA; WRKY transcription factor 1 | ||||
| STRING | GLYMA02G47650.1 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF9358 | 33 | 43 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G04880.1 | 5e-78 | zinc-dependent activator protein-1 | ||||



