 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | C.cajan_22366 | ||||||||
| Common Name | KK1_023024 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Cajanus
|
||||||||
| Family | TALE | ||||||||
| Protein Properties | Length: 390aa MW: 44959 Da PI: 6.9338 | ||||||||
| Description | TALE family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 28.6 | 2.4e-09 | 317 | 352 | 20 | 55 |
HHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHH CS
Homeobox 20 eknrypsaeereeLAkklgLterqVkvWFqNrRake 55
+k +yps++e+ LA+++gL+++q+ +WF N+R ++
C.cajan_22366 317 YKWPYPSESEKVALAESTGLDQKQINNWFINQRKRH 352
5679*****************************885 PP
| |||||||
| 2 | ELK | 35.4 | 2.3e-12 | 272 | 293 | 1 | 22 |
ELK 1 ELKhqLlrKYsgyLgsLkqEFs 22
ELK +LlrKYsgyL+sLkqE+s
C.cajan_22366 272 ELKTHLLRKYSGYLSSLKQELS 293
9*******************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01255 | 3.1E-19 | 128 | 172 | IPR005540 | KNOX1 |
| Pfam | PF03790 | 1.1E-20 | 130 | 171 | IPR005540 | KNOX1 |
| SMART | SM01256 | 4.3E-30 | 179 | 230 | IPR005541 | KNOX2 |
| Pfam | PF03791 | 1.3E-24 | 183 | 229 | IPR005541 | KNOX2 |
| Pfam | PF03789 | 3.5E-9 | 272 | 293 | IPR005539 | ELK domain |
| SMART | SM01188 | 4.8E-7 | 272 | 293 | IPR005539 | ELK domain |
| PROSITE profile | PS51213 | 11.051 | 272 | 292 | IPR005539 | ELK domain |
| PROSITE profile | PS50071 | 12.989 | 292 | 355 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 1.37E-19 | 293 | 368 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 2.1E-13 | 294 | 359 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 2.5E-27 | 297 | 358 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 2.99E-12 | 304 | 356 | No hit | No description |
| Pfam | PF05920 | 1.1E-16 | 312 | 351 | IPR008422 | Homeobox KN domain |
| PROSITE pattern | PS00027 | 0 | 330 | 353 | IPR017970 | Homeobox, conserved site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0001708 | Biological Process | cell fate specification | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010051 | Biological Process | xylem and phloem pattern formation | ||||
| GO:0010089 | Biological Process | xylem development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 390 aa Download sequence Send to blast |
MEEYTNNNNH LSENTGPRPS PRPNFLYSSS SGGNQNQNQS HHHHHHHHHQ QFPINTFHLQ 60 SGGSDHCFHE SDQVAPHPSV KTEATTSKLH APIFHYPLMR ANLHNNTMHH HHHHQGGASP 120 TSSTTEVEVI KAKIIAHPQY SNLLEAYMDC QKIGAPPEAV ARMVAAKQEF EARQRSSVGS 180 RETSKDPELD QFMEAYYDML VKYREELTRP IQEAMDFMRR IETQLNMLCN GPVRIFSDDK 240 CEGAGSSEEE DQDNSGGETE LPEIDPRAED RELKTHLLRK YSGYLSSLKQ ELSKKKKKGK 300 LPKDARQKLL NWWELHYKWP YPSESEKVAL AESTGLDQKQ INNWFINQRK RHWKPSEDMQ 360 FMVMDGLHPQ SATLYMDGHY MADGHYRLGP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probably binds to the DNA sequence 5'-TGAC-3'. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00675 | ChIP-seq | Transfer from GRMZM2G017087 | Download |
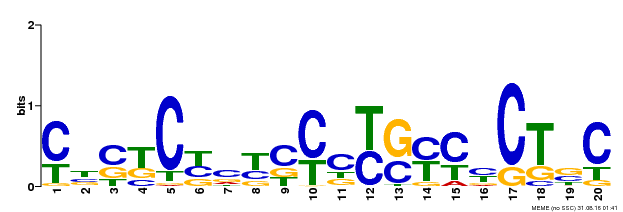
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | C.cajan_22366 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HM107002 | 0.0 | HM107002.1 Glycine max KNOX-like DNA-binding protein mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020222834.1 | 0.0 | homeotic protein knotted-1 isoform X1 | ||||
| Refseq | XP_029128578.1 | 0.0 | homeotic protein knotted-1 isoform X2 | ||||
| Swissprot | O04135 | 1e-176 | KNAP2_MALDO; Homeobox protein knotted-1-like 2 | ||||
| TrEMBL | A0A151T0Q6 | 0.0 | A0A151T0Q6_CAJCA; Homeobox protein knotted-1-like 2 | ||||
| STRING | GLYMA14G10430.1 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF4584 | 32 | 46 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G08150.1 | 1e-147 | KNOTTED-like from Arabidopsis thaliana | ||||



