 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | C.cajan_22309 | ||||||||
| Common Name | KK1_022967 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Cajanus
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1076aa MW: 121019 Da PI: 5.8137 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 180.5 | 2e-56 | 16 | 130 | 4 | 118 |
CG-1 4 ekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcy 99
++rwl++ ei++iL n++ +++t+e+++rp+sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+hekLKvg+v+vl+cyYah+een++fqrr+y
C.cajan_22309 16 AQHRWLRPAEICEILRNYRMFHITSEPHNRPPSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHEKLKVGSVDVLHCYYAHGEENENFQRRSY 111
49********************************************************************************************** PP
CG-1 100 wlLeeelekivlvhylevk 118
w+Le ++ +iv+vhylevk
C.cajan_22309 112 WMLEPDMMHIVFVHYLEVK 130
****************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 82.05 | 9 | 135 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 8.9E-81 | 12 | 130 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.9E-50 | 15 | 129 | IPR005559 | CG-1 DNA-binding domain |
| SuperFamily | SSF81296 | 1.91E-14 | 497 | 582 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF01833 | 8.5E-5 | 497 | 572 | IPR002909 | IPT domain |
| Gene3D | G3DSA:1.25.40.20 | 1.7E-19 | 680 | 792 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 19.969 | 680 | 791 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 1.12E-19 | 685 | 791 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 5.91E-16 | 686 | 789 | No hit | No description |
| SMART | SM00248 | 2200 | 697 | 726 | IPR002110 | Ankyrin repeat |
| Pfam | PF12796 | 6.4E-7 | 702 | 792 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 11.194 | 730 | 762 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 1.7E-4 | 730 | 759 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 2100 | 769 | 798 | IPR002110 | Ankyrin repeat |
| SMART | SM00015 | 0.14 | 899 | 921 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.316 | 900 | 929 | IPR000048 | IQ motif, EF-hand binding site |
| SuperFamily | SSF52540 | 2.44E-7 | 901 | 950 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| Pfam | PF00612 | 0.0026 | 902 | 920 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0034 | 922 | 944 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.432 | 923 | 947 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.3E-4 | 925 | 944 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1076 aa Download sequence Send to blast |
MPYYQCADIQ QLQFEAQHRW LRPAEICEIL RNYRMFHITS EPHNRPPSGS LFLFDRKVLR 60 YFRKDGHNWR KKKDGKTVKE AHEKLKVGSV DVLHCYYAHG EENENFQRRS YWMLEPDMMH 120 IVFVHYLEVK GNKNIVVNTE GDEVLSDSQK VTSPSSSFPT NHSGVPSLST DSVSPTSSLM 180 SLREDADSED IHQASSGLRP LHESQHMGNG PFTGKIDAGL NSSYPMHPLS GDHGQSTISG 240 VDYIPVPVAH GDKFRGNDTA YIDGHKTHGV APWDTVLEST AKFHNDPSLA SFPSITSSST 300 GNVLEQDHTI FGDLIMSKKG LTADAEISQS LQANWQIPFE DNSGDTPTLT QSFGLQFGSD 360 YGTGLLGNYT RNASSEIAPV LYNFHGESKE QTMQQNYPEE HADEQSQYAV KSNSANKIPD 420 EETINYGLNV KYALLDRDES LKKVDSFSRW VTKELGEVAD LNMQSSPGIS WSTDECQHVI 480 DDTSLSPSLS QDQLFSINDF SPKWAYAESE IEVLIIGSFL KSQPEVTTCK WSCMFGEVEV 540 PDEVLADGIL CCQAPRHKVG RVPFYVTCSN RLACSEVREF DFREGFARNV DFADFYNSST 600 EMLLHLRLED FLSLKPVDPS NHAFEGDLEK RNVIFKLISL REEEEYSIKE ESTTELDISQ 660 HKAKEHLFLR QVKEKLYSWL LHKVTESGKG PNILDEDGQG VLHLAAVLDY DWAINPIISA 720 GVNINFRDVN GWTALHWAAS YGRERTVAVL VSMGADCGAS TDPSPAFRSG RSAADLASSN 780 GHKGISGFLA ESSLTHHLET LTMDDQKSGM KVVQTVSERT ATPVIYGDMP DALCLKDSLT 840 AVRNATQAAD RIHQVYRMQS FQRKQLTQCE GDDQLGLSDQ QALSLLASRA GKSGQGDGLA 900 NAAATQIQKK FRGWKKRKEF LMIRQRIVKI QAHVRGHQVR KQYKTIIWSV GILEKVILRW 960 RRKGSGLRGF RPDAVNKVPS QQNDSPKEDD YDYLKEGRKQ KEEIIQKALS RVKSMAQYPE 1020 ARAQYRRLLN VVEDFRQTKA SDKGFLNSEE TVESVEDLID IDMLLDDDNF IPIAFD |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
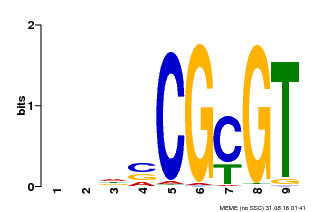
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | C.cajan_22309 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015038 | 0.0 | AP015038.1 Vigna angularis var. angularis DNA, chromosome 5, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020222227.1 | 0.0 | calmodulin-binding transcription activator 2 isoform X1 | ||||
| Refseq | XP_020222228.1 | 0.0 | calmodulin-binding transcription activator 2 isoform X2 | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A151T0H3 | 0.0 | A0A151T0H3_CAJCA; Calmodulin-binding transcription activator 2 | ||||
| STRING | GLYMA15G05900.2 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF7406 | 26 | 42 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



