 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | C.cajan_20664 | ||||||||
| Common Name | KK1_021281 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Cajanus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 687aa MW: 75750.1 Da PI: 6.7516 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 66.5 | 3.7e-21 | 43 | 98 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+k+l L++rqVk+WFqNrR+++k
C.cajan_20664 43 KKRYHRHTPQQIQELEALFKECPHPDEKQRLELSKRLCLETRQVKFWFQNRRTQMK 98
688999***********************************************999 PP
| |||||||
| 2 | START | 209.1 | 1.7e-65 | 204 | 428 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEEE CS
START 1 elaeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddkeqWdetla....kaetle 83
ela++a++elvk+a+ +ep+Wv+ + e++n +e++++f++ + + +ea r+ g+v+ ++ lve+l+d++ +W e+++ + +t+e
C.cajan_20664 204 ELALAAMDELVKMAQTGEPLWVRNVeggrEILNHEEYVRSFTPCIGlrpngFVSEATRENGMVIINSLALVETLMDSN-RWAEMFPciiaRTSTTE 298
5899**************************************9999****9***************************.***************** PP
EECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE- CS
START 84 vissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppesssvvRaellpSgiliepksnghskvtwvehvdl 172
vis+g galqlm aelq+lsplvp R++ f+R+++q+ +g+w++vdvS+ds ++ + + +v +++lpSg+++++++ng+skvtwveh+++
C.cajan_20664 299 VISNGingtrnGALQLMHAELQVLSPLVPvREVNFLRFCKQHAEGVWAVVDVSIDSIRESSGAPTFVNCRRLPSGCVVQDMPNGYSKVTWVEHAEY 394
***********************************************************999********************************** PP
-SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 173 kgrlphwllrslvksglaegaktwvatlqrqcek 206
+++++h+l+r+l++sg+ +g ++wvatlqrqce+
C.cajan_20664 395 DESQVHQLYRPLLSSGMGFGSQRWVATLQRQCEC 428
********************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 8.36E-21 | 26 | 100 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 9.2E-22 | 30 | 100 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.216 | 40 | 100 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 9.9E-18 | 41 | 104 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.89E-18 | 42 | 100 | No hit | No description |
| Pfam | PF00046 | 1.1E-18 | 43 | 98 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 75 | 98 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 44.574 | 195 | 431 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 4.4E-33 | 198 | 428 | No hit | No description |
| CDD | cd08875 | 7.14E-127 | 199 | 427 | No hit | No description |
| SMART | SM00234 | 4.1E-49 | 204 | 428 | IPR002913 | START domain |
| Pfam | PF01852 | 4.4E-57 | 204 | 428 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.1E-24 | 449 | 673 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 687 aa Download sequence Send to blast |
MDNFEPNGLR RSREEEHESR SGSDNMDGAS GDEHDAADNP PRKKRYHRHT PQQIQELEAL 60 FKECPHPDEK QRLELSKRLC LETRQVKFWF QNRRTQMKTQ LERHENTLLR QENDKLRAEN 120 MSIRDAMRNP MCSNCGGPAI IGEISLEEQH LRIENARLKD ELDRVCALAG KFLGRPVSSL 180 PNSSLELGVG GNNNGSVERS MFLELALAAM DELVKMAQTG EPLWVRNVEG GREILNHEEY 240 VRSFTPCIGL RPNGFVSEAT RENGMVIINS LALVETLMDS NRWAEMFPCI IARTSTTEVI 300 SNGINGTRNG ALQLMHAELQ VLSPLVPVRE VNFLRFCKQH AEGVWAVVDV SIDSIRESSG 360 APTFVNCRRL PSGCVVQDMP NGYSKVTWVE HAEYDESQVH QLYRPLLSSG MGFGSQRWVA 420 TLQRQCECLA ILMSSAAPSR DHSAITAGGR RSMLKLAQRM TNNFCAGVCA STVHKWNKLS 480 PGNVDEDVRV MTRKSPPGVV LSAATSVWLP VSPHRLFHFL RDERLRSEWD ILSNGGPMQE 540 MAHIAKGQDH GNCVSLLRAS AINSNQSSML ILQETCIDAA GSLVVYAPVD IPAMHVVMNG 600 GDSAYVALLP SGFAIVPDGP GSRGPTATTN GADNGGARVS GSLLTVAFQI LVNSLPTAKL 660 TVESVETVNN LISCTVQKIK AALHCES |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
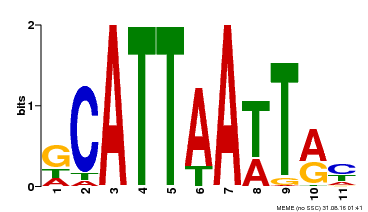
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | C.cajan_20664 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020237185.1 | 0.0 | homeobox-leucine zipper protein ANTHOCYANINLESS 2 isoform X1 | ||||
| Refseq | XP_020237261.1 | 0.0 | homeobox-leucine zipper protein ANTHOCYANINLESS 2 isoform X2 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A151UCG5 | 0.0 | A0A151UCG5_CAJCA; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| STRING | XP_007134961.1 | 0.0 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1192 | 34 | 106 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



