 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | C.cajan_17204 | ||||||||
| Common Name | KK1_017714 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Cajanus
|
||||||||
| Family | E2F/DP | ||||||||
| Protein Properties | Length: 500aa MW: 54874.6 Da PI: 4.9403 | ||||||||
| Description | E2F/DP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | E2F_TDP | 84.7 | 7.2e-27 | 182 | 247 | 1 | 71 |
E2F_TDP 1 rkeksLrlltqkflkllekseegivtlnevakeLvsedvknkrRRiYDilNVLealnliekkekneirwkg 71
r+++sL+llt+kf++l++++e+gi++ln++a++L +v ++RRiYDi+NVLe+++liekk kn+i+wkg
C.cajan_17204 182 RYDSSLGLLTKKFINLVKHAEDGILDLNKAAETL---EV--QKRRIYDITNVLEGIGLIEKKLKNRIHWKG 247
6899******************************...99..****************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.10 | 6.3E-29 | 179 | 249 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SMART | SM01372 | 1.2E-34 | 182 | 247 | IPR003316 | E2F/DP family, winged-helix DNA-binding domain |
| SuperFamily | SSF46785 | 7.35E-17 | 182 | 245 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| Pfam | PF02319 | 1.9E-24 | 184 | 247 | IPR003316 | E2F/DP family, winged-helix DNA-binding domain |
| CDD | cd14660 | 6.26E-48 | 260 | 364 | No hit | No description |
| SuperFamily | SSF144074 | 2.88E-32 | 260 | 364 | No hit | No description |
| Pfam | PF16421 | 2.6E-33 | 263 | 362 | IPR032198 | E2F transcription factor, CC-MB domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0051446 | Biological Process | positive regulation of meiotic cell cycle | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005667 | Cellular Component | transcription factor complex | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 500 aa Download sequence Send to blast |
MSSAAGAPDR RASPPQGAAG AASRPPLKRH LAFATKPPFA PPDEYHSFST PDSRRVADEA 60 VVVRSPFLLS LLLISFCLCS LSSRIWNVPF RLCGCDSLWF SYMKRKSGMN DTEGESQAQK 120 WSNSPGYTNV SNITSSPFKT PVSAKGGRTQ KAKASKEGRS CPPTPISNAG SPSPLTPASS 180 CRYDSSLGLL TKKFINLVKH AEDGILDLNK AAETLEVQKR RIYDITNVLE GIGLIEKKLK 240 NRIHWKGIES STSGEVDGDI SVLKAEVDKL TLEEQGLDDQ IREMQERLRN LSENENNQKC 300 LFVTEDDIKG LPCFQNETLI AIKAPHGTTL EVPDPEEAVD YPQRRYRIIL RSTMGPIDVY 360 LISQFEEKFE EINGAEHPMI PLASSSGSNE QLLTEMVPAD GKELETQTQL SSHAYSDLNA 420 SQEFAGGMMK IVPSDVDNDA DYWLLSDADV SITDMWRTDS TVDWNGVDIL HSDFGIVSRP 480 QTPSSGFAEV PPTGANSSQK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1cf7_A | 6e-25 | 177 | 248 | 2 | 74 | PROTEIN (TRANSCRIPTION FACTOR E2F-4) |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds DNA cooperatively with DP proteins through the E2 recognition site, 5'-TTTC[CG]CGC-3' found in the promoter region of a number of genes whose products are involved in cell cycle regulation or in DNA replication. The binding of retinoblastoma-related proteins represses transactivation. Regulates gene expression both positively and negatively. Activates the expression of E2FB. Involved in the control of cell-cycle progression from G1 to S phase. Stimulates cell proliferation and delays differentiation. {ECO:0000269|PubMed:11669580, ECO:0000269|PubMed:11786543, ECO:0000269|PubMed:11862494, ECO:0000269|PubMed:11889041, ECO:0000269|PubMed:11891240, ECO:0000269|PubMed:12913157, ECO:0000269|PubMed:15377755, ECO:0000269|PubMed:16514015, ECO:0000269|PubMed:19662336}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00293 | DAP | Transfer from AT2G36010 | Download |
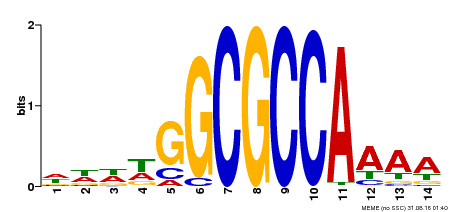
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | C.cajan_17204 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT093046 | 1e-108 | BT093046.1 Soybean clone JCVI-FLGm-15I8 unknown mRNA. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020221307.1 | 0.0 | transcription factor E2FA isoform X2 | ||||
| Swissprot | Q9FNY0 | 1e-155 | E2FA_ARATH; Transcription factor E2FA | ||||
| TrEMBL | A0A151T7V5 | 0.0 | A0A151T7V5_CAJCA; Transcription factor E2F3 | ||||
| STRING | GLYMA02G35850.1 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1947 | 34 | 90 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G36010.1 | 1e-147 | E2F transcription factor 3 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



