 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | C.cajan_00424 | ||||||||
| Common Name | KK1_000439 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Cajanus
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 837aa MW: 92256.7 Da PI: 6.4632 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 57.7 | 2e-18 | 16 | 74 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++ +++ps +r++L +++ +++ +q+kvWFqNrR +ek+
C.cajan_00424 16 KYVRYTPEQVEALERLYHDCPKPSSIRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 74
5679*****************************************************97 PP
| |||||||
| 2 | START | 178.4 | 4.1e-56 | 160 | 368 | 2 | 205 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECTT..EEEEEEE CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlevissg..galqlmv 95
+aee+++e+++ka+ ++ Wv+++ +++g++++ +++ s++++g a+ra+g+v +++ v+e+l+d++ W ++++++++l+v+ ++ g+++l +
C.cajan_00424 160 IAEETLAEFLSKATGTAVEWVQMPGMKPGPDSIGIVAISHGCTGVAARACGLVGLEPT-RVAEILKDRPSWFRDCRAVDVLNVLPTAngGTIELLY 254
7899******************************************************.8888888888****************9999******* PP
EXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHHHHHHHHHH CS
START 96 aelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwvehvdlkgrlphwllrslvks 187
++l+a+++l+p Rdf+ +Ry+ l++g++vi+++S+ ++q+ p+ +++vRae+lpSg+li+p+++g+s +++v+h+dl+ ++++++lr+l++s
C.cajan_00424 255 MQLYAPTTLAPaRDFWLLRYTSVLEDGSLVICERSLKNTQNGPSmppVQHFVRAEMLPSGYLIRPCEGGGSIIHIVDHMDLEPWSVPEVLRPLYES 350
******************************************9988899*********************************************** PP
HHHHHHHHHHHHTXXXXX CS
START 188 glaegaktwvatlqrqce 205
+++ ++kt++ +l+++++
C.cajan_00424 351 STVLAQKTTMVALRQLRQ 368
***********9999876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.402 | 11 | 75 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.2E-15 | 13 | 79 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 6.42E-17 | 14 | 79 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 1.31E-16 | 16 | 76 | No hit | No description |
| Pfam | PF00046 | 5.8E-16 | 17 | 74 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 3.5E-18 | 18 | 74 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 1.27E-6 | 68 | 107 | No hit | No description |
| PROSITE profile | PS50848 | 25.241 | 150 | 365 | IPR002913 | START domain |
| CDD | cd08875 | 9.62E-80 | 154 | 369 | No hit | No description |
| Gene3D | G3DSA:3.30.530.20 | 2.4E-23 | 157 | 365 | IPR023393 | START-like domain |
| SMART | SM00234 | 8.9E-41 | 159 | 369 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 8.52E-38 | 159 | 348 | No hit | No description |
| Pfam | PF01852 | 1.5E-53 | 160 | 368 | IPR002913 | START domain |
| Pfam | PF08670 | 1.2E-51 | 694 | 836 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 837 aa Download sequence Send to blast |
MAMSCKDGKH GLDNGKYVRY TPEQVEALER LYHDCPKPSS IRRQQLIREC PILSNIEPKQ 60 IKVWFQNRRC REKQRKEASR LQAVNRKLTA MNKLLMEEND RLQKQVSQLV YENGYFRQHT 120 QNTALPTKDT SCESAVTSGQ RSLTTQHPPR DASPAGLMSI AEETLAEFLS KATGTAVEWV 180 QMPGMKPGPD SIGIVAISHG CTGVAARACG LVGLEPTRVA EILKDRPSWF RDCRAVDVLN 240 VLPTANGGTI ELLYMQLYAP TTLAPARDFW LLRYTSVLED GSLVICERSL KNTQNGPSMP 300 PVQHFVRAEM LPSGYLIRPC EGGGSIIHIV DHMDLEPWSV PEVLRPLYES STVLAQKTTM 360 VALRQLRQIS HEVSQSNVTG WGRRPAALRA LGQRLSRGFN EAINGFTDEG WSMIGNDGVD 420 DVTVLVNSAP DKLMGLNLSF ANGFPAVSNA VLCAKASMLL QNVPPAILLR FLREHRSEWA 480 DNNMDAYSAA AIKVGPCGLP GSRVGNYGGQ VILPLAHTIE HEEFLEVIKL EGIASSPEET 540 IMPREVFLLQ LCSGMDENAV GTCAELIFAP IDASFADDAP LLPSGFRIIP LDSGKEASNP 600 NRTLDLASAL DVGPTGNRAS NDYSGNSGCM RSVMTIAFEF AFESHMQEHV ASMARQYVRS 660 IISSVQRVAL ALSPSHLSSQ AGLRTPLGTP EAQTLARWIC NSYRCYLGVE LLKSNNEGNE 720 SLLKSLWHHS DAILCCTLKA LPVFTFANQA GLDMLETTLV ALQDITLEKI FDDHGRKILC 780 SEFPQIIQQG FACLQGGICL SSMGRPISYE RVVAWKVLNE DENAHCICFM FMNWSFV |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
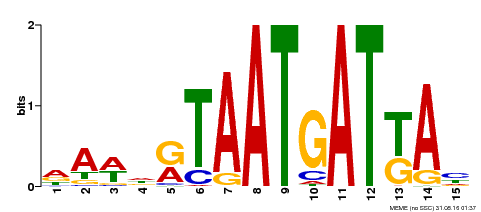
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | C.cajan_00424 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020226055.1 | 0.0 | homeobox-leucine zipper protein ATHB-15 | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | A0A151SHL8 | 0.0 | A0A151SHL8_CAJCA; Homeobox-leucine zipper protein ATHB-15 | ||||
| STRING | XP_007150614.1 | 0.0 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1946 | 33 | 87 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



