 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Csa18g035470.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Camelina
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 240aa MW: 26947.6 Da PI: 5.8233 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 43.3 | 8.3e-14 | 17 | 67 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT...-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGgg...tWktIartmgkgRtlkqcksrwqkyl 48
+g+WT eEd +lvd + G+g +W+ +++ g++R++k+c++rw +yl
Csa18g035470.1 17 KGPWTVEEDGKLVDFLRIRGSGggwCWRDVPKLAGLRRCGKSCRLRWTNYL 67
79***********************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 55.2 | 1.6e-17 | 73 | 117 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
rg +++eE +l++d++++lG++ W++Ia +++ gRt++++k++w+++
Csa18g035470.1 73 RGLFSEEEIQLIIDLHARLGNR-WSKIAVELP-GRTDNDIKNYWNTH 117
7889******************.*********.************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 5.0E-18 | 8 | 69 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 12.477 | 12 | 67 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 7.11E-25 | 14 | 114 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 7.4E-7 | 16 | 69 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.9E-12 | 17 | 67 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.76E-5 | 19 | 67 | No hit | No description |
| PROSITE profile | PS51294 | 26.005 | 68 | 122 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 7.2E-26 | 70 | 123 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 7.7E-17 | 72 | 120 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 7.6E-16 | 73 | 117 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.57E-12 | 76 | 118 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 240 aa Download sequence Send to blast |
MIMGGRKPCC DEVGLRKGPW TVEEDGKLVD FLRIRGSGGG WCWRDVPKLA GLRRCGKSCR 60 LRWTNYLRPD LKRGLFSEEE IQLIIDLHAR LGNRWSKIAV ELPGRTDNDI KNYWNTHIKR 120 KLIRMGIDPN THRRFDQQKV NEEEKVLKVN SPKPPSQPEV VVPVVLQNDT SAGNLNELAD 180 VDGDDQPWSF LMENDGGGAG ELTMLLSGNI TSSCSSSSSL WLKYGDLGYE DLELGCFDD* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1h8a_C | 5e-27 | 15 | 122 | 25 | 128 | MYB TRANSFORMING PROTEIN |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that acts as positive regulator of abscisic acid (ABA) signaling in response to salt stress. Acts as negative regulator ABI1, ABI2 and PP2CA, which are protein phosphatases 2C acting as negative regulator of ABA signaling. Binds to the DNA specific sequence and core element 5'-ACGT-3' found in the promoters of ABI1 and PP2CA to negatively regulate their expression during ABA-dependent salt stress response. {ECO:0000269|PubMed:23660402}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00577 | DAP | Transfer from AT5G62320 | Download |
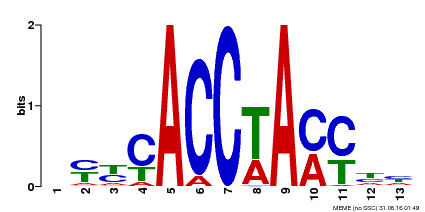
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Csa18g035470.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by salt stress, drought stress and abscisic acid (ABA). Down-regulated by salicylic acid (SA) methyl jasmonate (JA). {ECO:0000269|PubMed:23660402}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF199026 | 0.0 | AF199026.2 Arabidopsis thaliana putative transcription factor (MYB99) mRNA, complete cds. | |||
| GenBank | AY519644 | 0.0 | AY519644.1 Arabidopsis thaliana MYB transcription factor (At5g62320) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010483899.1 | 1e-176 | PREDICTED: protein ODORANT1-like | ||||
| Swissprot | Q9C7U7 | 1e-67 | MYB20_ARATH; Transcription factor MYB20 | ||||
| TrEMBL | R0F0I2 | 1e-143 | R0F0I2_9BRAS; Uncharacterized protein | ||||
| STRING | XP_010483899.1 | 1e-175 | (Camelina sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4 | 28 | 2646 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62320.1 | 1e-129 | myb domain protein 99 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



