 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Csa15g001260.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Camelina
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 443aa MW: 51253.4 Da PI: 6.3329 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 53.6 | 3.7e-17 | 74 | 125 | 5 | 56 |
SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 5 ttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ eq+++Le+ Fe +++ e++ +LAk lg++ rq+ +WFqNrRa++k
Csa15g001260.1 74 KRLQLEQVKALEKSFELGNKLEPERKIQLAKALGMQPRQIAIWFQNRRARWK 125
457889*********************************************9 PP
| |||||||
| 2 | Homeobox | 53.6 | 3.7e-17 | 244 | 295 | 5 | 56 |
SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 5 ttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ eq+++Le+ Fe +++ e++ +LAk lg++ rq+ +WFqNrRa++k
Csa15g001260.1 244 KRLQLEQVKALEKSFELGNKLEPERKIQLAKALGMQPRQIAIWFQNRRARWK 295
457889*********************************************9 PP
| |||||||
| 3 | HD-ZIP_I/II | 122.1 | 2.8e-39 | 71 | 161 | 1 | 91 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreel 91
ekk+rl+ eqvk+LE+sFe +kLeperK +la++Lg+qprq+a+WFqnrRAR+kt+qlE+dy++Lk+++++lk++n++L + +++L +e+
Csa15g001260.1 71 EKKKRLQLEQVKALEKSFELGNKLEPERKIQLAKALGMQPRQIAIWFQNRRARWKTRQLERDYDSLKKQFESLKSDNDSLLAYNKKLLAEV 161
69************************************************************************************99887 PP
| |||||||
| 4 | HD-ZIP_I/II | 122 | 3e-39 | 241 | 332 | 1 | 92 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreelk 92
ekk+rl+ eqvk+LE+sFe +kLeperK +la++Lg+qprq+a+WFqnrRAR+kt+qlE+dy++Lk+++++lk++n++L + +++L +e++
Csa15g001260.1 241 EKKKRLQLEQVKALEKSFELGNKLEPERKIQLAKALGMQPRQIAIWFQNRRARWKTRQLERDYDSLKKQFESLKSDNDSLLAYNKKLLAEVM 332
69*************************************************************************************99986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 16.115 | 67 | 127 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 4.6E-16 | 70 | 131 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 5.56E-18 | 70 | 129 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 4.21E-14 | 72 | 128 | No hit | No description |
| Pfam | PF00046 | 2.3E-14 | 73 | 125 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 6.8E-18 | 75 | 135 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 6.4E-5 | 98 | 107 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 102 | 125 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 6.4E-5 | 107 | 123 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 2.1E-11 | 128 | 161 | IPR003106 | Leucine zipper, homeobox-associated |
| PROSITE profile | PS50071 | 16.115 | 237 | 297 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 5.56E-18 | 240 | 299 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 4.6E-16 | 240 | 301 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 4.21E-14 | 242 | 298 | No hit | No description |
| Pfam | PF00046 | 2.3E-14 | 243 | 295 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 6.8E-18 | 245 | 305 | IPR009057 | Homeodomain-like |
| PROSITE pattern | PS00027 | 0 | 272 | 295 | IPR017970 | Homeobox, conserved site |
| Pfam | PF02183 | 9.2E-14 | 298 | 337 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 443 aa Download sequence Send to blast |
MAFPQHGFMF QQLHEDTSQD QLPSCPPHLF SGGGNYMMNR SMSLMNVQED HNQTLDEENL 60 SDDGAHTMLG EKKKRLQLEQ VKALEKSFEL GNKLEPERKI QLAKALGMQP RQIAIWFQNR 120 RARWKTRQLE RDYDSLKKQF ESLKSDNDSL LAYNKKLLAE VYADAGLRLE MAFPQHGFMF 180 QQLHEDTSQD QLPSCPPHLF SGGGNYMMNR SMSLMNVQED HNQTLDEENL SDDGAHTMLG 240 EKKKRLQLEQ VKALEKSFEL GNKLEPERKI QLAKALGMQP RQIAIWFQNR RARWKTRQLE 300 RDYDSLKKQF ESLKSDNDSL LAYNKKLLAE VMALKNKECN ESNIIKREAE ASWSNNGSTE 360 NSSDINLEMP REITTTHVNT IKDLFPSSIR SSAHRDNDDH HQNHEMVQEE SLCNMFNGID 420 ETTSAGYWAW SDPNHNHPHQ FN* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
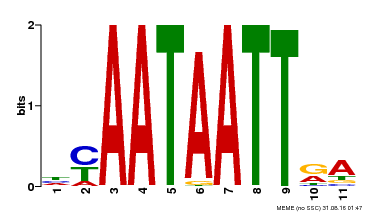
| Motif ID | Method | Source | Motif file |
| MP00322 | DAP | Transfer from AT3G01220 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Csa15g001260.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. {ECO:0000269|PubMed:12644682}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK228890 | 0.0 | AK228890.1 Arabidopsis thaliana mRNA for putative homeobox-leucine zipper protein, complete cds, clone: RAFL16-20-K05. | |||
| GenBank | AY087631 | 0.0 | AY087631.1 Arabidopsis thaliana clone 3722 mRNA, complete sequence. | |||
| GenBank | BT029221 | 0.0 | BT029221.1 Arabidopsis thaliana At3g01220 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010463554.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-20 isoform X1 | ||||
| Refseq | XP_010463555.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-20 isoform X2 | ||||
| Swissprot | Q8LAT0 | 0.0 | ATB20_ARATH; Homeobox-leucine zipper protein ATHB-20 | ||||
| TrEMBL | D7LAH8 | 0.0 | D7LAH8_ARALL; ATHB20 | ||||
| STRING | XP_010463555.1 | 0.0 | (Camelina sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3095 | 26 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G01220.1 | 0.0 | homeobox protein 20 | ||||



