 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Csa13g012280.2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Camelina
|
||||||||
| Family | AP2 | ||||||||
| Protein Properties | Length: 591aa MW: 65914 Da PI: 6.8005 | ||||||||
| Description | AP2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 31.4 | 4.6e-10 | 281 | 337 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrd.pseng...krkrfslgkfgtaeeAakaaiaarkkleg 55
s y+GV++++++gr++A+++d + rk + g ++ +++Aa+a++ a++k+++
Csa13g012280.2 281 SIYRGVTRHRWTGRYEAHLWDnSCR-RegqARKGRQ-GGYDKEDKAARAYDLAALKYWN 337
57*******************4444.2344336655.7799999*************97 PP
| |||||||
| 2 | AP2 | 49.6 | 9.8e-16 | 380 | 431 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s y+GV++++++grW A+I + +k +lg+f t+eeAa+a++ a+ k++g
Csa13g012280.2 380 SIYRGVTRHHQQGRWQARIGRVAG---NKDLYLGTFATEEEAAEAYDIAAIKFRG 431
57****************988532...5************************997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF00847 | 7.8E-8 | 281 | 337 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 1.63E-13 | 281 | 346 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 2.36E-17 | 281 | 347 | No hit | No description |
| SMART | SM00380 | 1.8E-18 | 282 | 351 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 16.753 | 282 | 345 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 1.3E-10 | 282 | 345 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 5.4E-6 | 283 | 294 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 3.51E-11 | 380 | 439 | No hit | No description |
| SuperFamily | SSF54171 | 1.11E-17 | 380 | 440 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 4.1E-10 | 380 | 431 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 7.9E-31 | 381 | 445 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.111 | 381 | 439 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 5.6E-18 | 381 | 439 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 5.4E-6 | 421 | 441 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010080 | Biological Process | regulation of floral meristem growth | ||||
| GO:0010492 | Biological Process | maintenance of shoot apical meristem identity | ||||
| GO:0035265 | Biological Process | organ growth | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:0060772 | Biological Process | leaf phyllotactic patterning | ||||
| GO:0060774 | Biological Process | auxin mediated signaling pathway involved in phyllotactic patterning | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 591 aa Download sequence Send to blast |
MAPMTNWLTF SLSPMEMLRS SDQSQFVSYD ASSAASSSPY LLDNFYGWTN QKPQEFFKDE 60 AQLAVAASMA DSTILTTFVD PQSHSQSHIP KLEDFLGDSS SMVRYSDNSQ TETQDSSLTQ 120 IYDPRHHQNH HHNQTGFYSD HHDFKTMAGF QTAFSTNSGS EVDDSASIGR THLAGEYLGH 180 VVESSSGPDL GFHGGSNNTT NTGGALSLGV NVNNSTNQHR NDNEITNHHY RGGGERNNNN 240 NNNNNENNEK TESEKEKPIV AVETSDCSNK KIADTFGQRT SIYRGVTRHR WTGRYEAHLW 300 DNSCRREGQA RKGRQGGYDK EDKAARAYDL AALKYWNATA TTNFPITNYS KELEDMKHMT 360 KQEFIASLRR KSSGFSRGAS IYRGVTRHHQ QGRWQARIGR VAGNKDLYLG TFATEEEAAE 420 AYDIAAIKFR GINAVTNFEM NRYDVEAIMK SALPIGGAAK RLKLSLEAAS SEQKPILGHQ 480 LHHFQQQQQQ QQQLQLQSSP NHSSINFALC PNPAVQSQQI IPCGIPFEAA ALYHHQQQQQ 540 QQQQNFFQHF PANAASDSTG SNNNSNVQGS MGLMAPNPAE FFLWPNNQSY * |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probably acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. May be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways (By similarity). {ECO:0000250}. | |||||
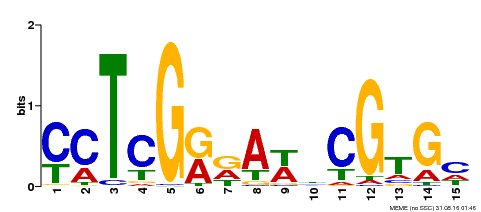
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00504 | DAP | Transfer from AT5G10510 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Csa13g012280.2 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY974197 | 0.0 | AY974197.1 Arabidopsis thaliana clone At5g10510 AP2/EREBP transcription factor mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010453080.1 | 0.0 | PREDICTED: AP2-like ethylene-responsive transcription factor AIL6 | ||||
| Swissprot | Q52QU2 | 0.0 | AIL6_ARATH; AP2-like ethylene-responsive transcription factor AIL6 | ||||
| TrEMBL | D7M2X9 | 0.0 | D7M2X9_ARALL; Uncharacterized protein | ||||
| STRING | XP_010453080.1 | 0.0 | (Camelina sativa) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G10510.2 | 0.0 | AINTEGUMENTA-like 6 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



