 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Csa12g017140.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Camelina
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 676aa MW: 74647.4 Da PI: 6.9667 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 77.3 | 1.7e-24 | 122 | 223 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSS CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrs 88
f k+lt+sd+++ g +++p+ +ae++ +++ + t+ +d +g +W++++iyr++++r++lt+GW++Fv++++L +gD++vF+ + +
Csa12g017140.1 122 FAKTLTQSDANNGGGFSVPRYCAETIfprldynAEPPVQ--TILAKDVHGDVWKFRHIYRGTPRRHLLTTGWSNFVNQKKLVAGDSIVFM--RAE 212
89****************************985444444..8999*********************************************..448 PP
EE..EEEEE-S CS
B3 89 efelvvkvfrk 99
++ l+v+++r+
Csa12g017140.1 213 NGDLCVGIRRA 223
8999****997 PP
| |||||||
| 2 | Auxin_resp | 113.7 | 1.7e-37 | 281 | 364 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrl.sGtvvgvsdldpvrWpnSkWrsLk 83
aa+ a ++ +FevvY+PrastseF+vk+ + ++a++++++ GmRfkmafetedss++++ +Gtv++v+ +dp+rWpnS+Wr+L+
Csa12g017140.1 281 AATLAISGRPFEVVYYPRASTSEFCVKALDARAAMRIPWCSGMRFKMAFETEDSSRISWfMGTVSAVNVSDPIRWPNSPWRLLQ 364
67788899**************************************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 5.6E-39 | 114 | 229 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 7.85E-30 | 117 | 225 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 1.71E-22 | 120 | 222 | No hit | No description |
| PROSITE profile | PS50863 | 13.352 | 122 | 224 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 1.8E-22 | 122 | 223 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 7.1E-23 | 122 | 224 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 2.7E-31 | 281 | 364 | IPR010525 | Auxin response factor |
| PROSITE profile | PS51745 | 18.27 | 589 | 669 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007389 | Biological Process | pattern specification process | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0048829 | Biological Process | root cap development | ||||
| GO:0051301 | Biological Process | cell division | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 676 aa Download sequence Send to blast |
MINVMNPMKS GSEKGLDPQL WHACAGGMVR MPPMNSKVFY FPQGHAENAY DCVDFGNLPI 60 HPMVLCRVLA IKYMADPESD EVFAKLRLIP LKEDEYVDHH HEYADGEDSN GFESNSEKTP 120 SFAKTLTQSD ANNGGGFSVP RYCAETIFPR LDYNAEPPVQ TILAKDVHGD VWKFRHIYRG 180 TPRRHLLTTG WSNFVNQKKL VAGDSIVFMR AENGDLCVGI RRAKRGGIGN GPEYSAGWNP 240 IGGSCGYSSL LREDESNSLR RSNCSLADRK GKVTAESVIE AATLAISGRP FEVVYYPRAS 300 TSEFCVKALD ARAAMRIPWC SGMRFKMAFE TEDSSRISWF MGTVSAVNVS DPIRWPNSPW 360 RLLQVAWDEP DLLQNVKRVN PWLVELVSNV HPIPLTSFSP PRKKMRLPQH PDYNNLINSI 420 PIPSFPSNPL IRSSPLSSVL DNVPVGLQGA RHNAHQYYGL SSSDLHHYYL NRQPPPPPPP 480 SSLPLSPSLG LRNIDTKNEK GFCFLTMGTT PCNDTESKKS SHIVLFGKLI LPEEQLSEKG 540 STDTANLEKT QISSGGSNQN NGIAGRDLSS SDEGSPCSKK VQDASGLETG HCKVFMESDD 600 VGRTLDLSVL GSYEELSRKL SDMFGIQKSE MLSSVLYRDA SGAIKYAGNE PFSEFLKTAR 660 RLTILTEQGS ESVVI* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldu_A | 5e-76 | 9 | 385 | 43 | 388 | Auxin response factor 5 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). Could act as transcriptional activator or repressor. Formation of heterodimers with Aux/IAA proteins may alter their ability to modulate early auxin response genes expression. {ECO:0000269|PubMed:12036261}. | |||||
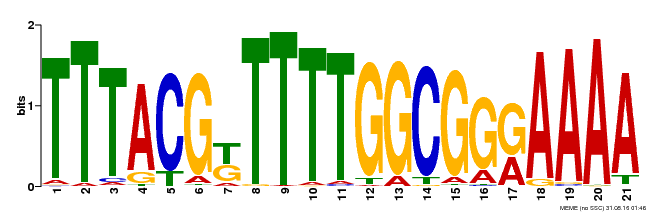
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00461 | DAP | Transfer from AT4G30080 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Csa12g017140.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY059792 | 0.0 | AY059792.1 Arabidopsis thaliana putative transcription factor (At4g30080) mRNA, complete cds. | |||
| GenBank | AY091198 | 0.0 | AY091198.1 Arabidopsis thaliana putative transcription factor (At4g30080) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010447772.1 | 0.0 | PREDICTED: auxin response factor 16 | ||||
| Swissprot | Q93YR9 | 0.0 | ARFP_ARATH; Auxin response factor 16 | ||||
| TrEMBL | R0GY89 | 0.0 | R0GY89_9BRAS; Auxin response factor | ||||
| STRING | XP_010447772.1 | 0.0 | (Camelina sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM580 | 27 | 130 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G30080.1 | 0.0 | auxin response factor 16 | ||||



