 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Csa10g019030.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Camelina
|
||||||||
| Family | Nin-like | ||||||||
| Protein Properties | Length: 965aa MW: 106107 Da PI: 5.8856 | ||||||||
| Description | Nin-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | RWP-RK | 91.8 | 5e-29 | 595 | 645 | 2 | 52 |
RWP-RK 2 ekeisledlskyFslpikdAAkeLgvclTvLKriCRqyGIkRWPhRkiksl 52
ek+isl++l++yF +++kdAAk+Lgvc+T++KriCRq+GI+RWP+Rkik++
Csa10g019030.1 595 EKTISLDVLQQYFTGSLKDAAKSLGVCPTTMKRICRQHGISRWPSRKIKKV 645
799**********************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF55781 | 3.63E-5 | 218 | 289 | IPR029016 | GAF domain-like |
| PROSITE profile | PS51519 | 17.757 | 584 | 665 | IPR003035 | RWP-RK domain |
| Pfam | PF02042 | 7.4E-26 | 597 | 645 | IPR003035 | RWP-RK domain |
| SuperFamily | SSF54277 | 1.86E-23 | 864 | 953 | No hit | No description |
| Pfam | PF00564 | 3.1E-20 | 868 | 949 | IPR000270 | PB1 domain |
| PROSITE profile | PS51745 | 27.975 | 868 | 950 | IPR000270 | PB1 domain |
| Gene3D | G3DSA:3.10.20.240 | 2.5E-27 | 868 | 950 | No hit | No description |
| SMART | SM00666 | 2.1E-28 | 868 | 950 | IPR000270 | PB1 domain |
| CDD | cd06407 | 1.71E-34 | 869 | 949 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0010167 | Biological Process | response to nitrate | ||||
| GO:0042128 | Biological Process | nitrate assimilation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 965 aa Download sequence Send to blast |
MCEPDDNSGR NGVAHPPSSR SRELLMDVDD LDLDGSWPLD QIPYLSSSNR MISPIFVSSS 60 SEQPCSPLWA FSDGGGNGGI HHHANGGGGD EEKNSSASGV PSSFRLAEYP LFLPYSSPSA 120 AENTTEKHNN NFQFPSPLMS LVPPENADNY CVIKERMTQA LRYFKDSTEQ HVLAQVWAPV 180 RKNGRDLLTT LGQPFVLNPN GNGLNQYRMI SLTYMFSVDS ESDIELGLPG RVFRQKLPEW 240 TPNVQYYSSK EFSRLDHALH YNVRGTLALP VFNPSGQSCI GVVELIMTSE KIHYAPEVDK 300 VCKALEAVNL KSSEILDHQT TQICNESRQN ALAEILEVLT VVCETHNLPL AQTWVPCQHG 360 SVLANGGGLK KNCTSFDGSC MGQICMSTTD MACYVVDAHV WGFRDACLEH HLQKGQGVAG 420 RAFLNGGSCF CRDITKFCKT QYPLVHYALM FKLTTCFAIS LQSSYTGDDS YILEFFLPSS 480 ITDDQEQDSL LGSILVTMKE HFQSLRVASG VDFGEDDDKL SFEIIQALPD KKIHSKIESI 540 RVPFSGSKSN AIETRLIPQP AVQSSDPVKE NINVATVNGV VKEKKKTEKK RGKTEKTISL 600 DVLQQYFTGS LKDAAKSLGV CPTTMKRICR QHGISRWPSR KIKKVNRSIT KLKRVIESVQ 660 GTDGGLDLTS MAVSSIPWTH GQTSAQPLNS PSGSKPPEVP NTNNSPNHWS SDHSPHEPNG 720 SPELPSNGHK RSRTVDESAG TPTSHGSCDG NQLDEAKVPN QDPLFTVGES PGLFFPPYSR 780 DHDVSAASFS MPNRLLGSID HFRGMLIEDA GSSKDLRNLC STTAFDDKFP DSNWMNNDNN 840 SNNNVYAPPK EEAIANVTRE PSGSETRTVT IKASYKDDII RFRISSGSGI MELKDEVAKR 900 LKLDAGTFDI KYLDDDNEWV LIACDADLQE CLEIPRSSRT NIVRLLVHDV TTNLGSSCES 960 TGEL* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in regulation of nitrate assimilation and in transduction of the nitrate signal. {ECO:0000269|PubMed:18826430}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
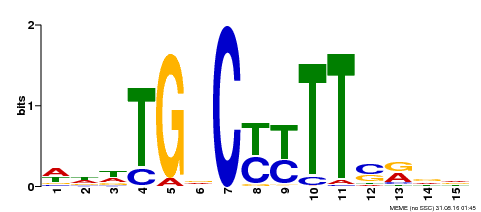
| Motif ID | Method | Source | Motif file |
| MP00449 | DAP | Transfer from AT4G24020 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Csa10g019030.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Not regulated by the N source or by nitrate. {ECO:0000269|PubMed:18826430}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK228452 | 0.0 | AK228452.1 Arabidopsis thaliana mRNA for hypothetical protein, complete cds, clone: RAFL15-03-P13. | |||
| GenBank | BT005776 | 0.0 | BT005776.1 Arabidopsis thaliana clone RAFL15-03-P13 (R20174) unknown protein (At4g24020) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010433823.1 | 0.0 | PREDICTED: protein NLP7 isoform X1 | ||||
| Swissprot | Q84TH9 | 0.0 | NLP7_ARATH; Protein NLP7 | ||||
| TrEMBL | R0GSX5 | 0.0 | R0GSX5_9BRAS; Uncharacterized protein | ||||
| STRING | XP_010433823.1 | 0.0 | (Camelina sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2507 | 26 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G24020.1 | 0.0 | NIN like protein 7 | ||||



