 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Csa03g060450.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Camelina
|
||||||||
| Family | TCP | ||||||||
| Protein Properties | Length: 396aa MW: 43355.4 Da PI: 6.8671 | ||||||||
| Description | TCP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCP | 117.9 | 1.3e-36 | 48 | 155 | 2 | 78 |
TCP 2 agkkdrhskihTkvggRdRRvRlsaecaarfFdLqdeLGfdkdsktieWLlqqakpaikeltgt.............................ss 67
+g+kdrhsk++T++g+RdRRvRlsa++a++f+d+qd+LGfd++sk+++WL+++ak ai++l+++ ++
Csa03g060450.1 48 TGRKDRHSKVCTAKGPRDRRVRLSAPTAIQFYDVQDRLGFDRPSKAVDWLIKKAKSAIDDLAELppwnpadalrqhaanakprktktlvspppPQ 142
79**************************************************************6555555554444444444444443333211 PP
TCP 68 ssasec..eaess 78
+++++ e++++
Csa03g060450.1 143 PPPQHQetEHHHH 155
1111111111111 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03634 | 5.3E-33 | 48 | 156 | IPR005333 | Transcription factor, TCP |
| PROSITE profile | PS51369 | 34.272 | 50 | 108 | IPR017887 | Transcription factor TCP subgroup |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0030154 | Biological Process | cell differentiation | ||||
| GO:0045962 | Biological Process | positive regulation of development, heterochronic | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 396 aa Download sequence Send to blast |
MAANNHLVDS PSPPMRHNQS AFTASPTENG DGCGEIVEVQ GGHIVRSTGR KDRHSKVCTA 60 KGPRDRRVRL SAPTAIQFYD VQDRLGFDRP SKAVDWLIKK AKSAIDDLAE LPPWNPADAL 120 RQHAANAKPR KTKTLVSPPP PQPPPQHQET EHHHHQIGEE DNESSFLPAS MDSDSIADTI 180 KSFFPVASSQ QSYHHHQPPS RAANTQNQDL LRLSLQSFQN GPPFTDQTEP VLFSGQSNHP 240 LGFDSSTAGW DQNHQSPEFG KIQRLVTWNN GGAADSAGST GGEGYVFASP SPVFSQSQLL 300 SQRGPLQSIN NTPMIRAWFD PHHHHQSMTT DDLHHHHHHP YHVPTGIHQS AIPGIAFASS 360 GEFSGFRIPA RFHGEQEDEH GGDNKPSSAS SDSRH* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5zkt_A | 4e-20 | 55 | 108 | 1 | 54 | Putative transcription factor PCF6 |
| 5zkt_B | 4e-20 | 55 | 108 | 1 | 54 | Putative transcription factor PCF6 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Plays a pivotal role in the control of morphogenesis of shoot organs by negatively regulating the expression of boundary-specific genes such as CUC genes, probably through the induction of miRNA (e.g. miR164). Participates in ovule develpment (PubMed:25378179). {ECO:0000269|PubMed:17307931, ECO:0000269|PubMed:25378179}. | |||||
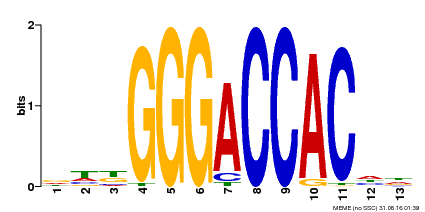
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00203 | DAP | Transfer from AT1G53230 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Csa03g060450.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC008007 | 0.0 | AC008007.7 Genomic sequence for Arabidopsis thaliana BAC F12M16 from chromosome I, complete sequence. | |||
| GenBank | AY099678 | 0.0 | AY099678.1 Arabidopsis thaliana putative flower development protein cycloidea (At1g53230) mRNA, complete cds. | |||
| GenBank | BT000266 | 0.0 | BT000266.1 Arabidopsis thaliana putative flower development protein cycloidea (At1g53230) mRNA, complete cds. | |||
| GenBank | CP002684 | 0.0 | CP002684.1 Arabidopsis thaliana chromosome 1 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010500959.1 | 0.0 | PREDICTED: transcription factor TCP3-like | ||||
| Swissprot | Q9MAH8 | 0.0 | TCP3_ARATH; Transcription factor TCP3 | ||||
| TrEMBL | D7KKF4 | 0.0 | D7KKF4_ARALL; Uncharacterized protein | ||||
| STRING | XP_010500959.1 | 0.0 | (Camelina sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2408 | 27 | 71 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G53230.1 | 0.0 | TEOSINTE BRANCHED 1, cycloidea and PCF transcription factor 3 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



