 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Csa02g070050.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Camelina
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 860aa MW: 95669.7 Da PI: 6.4454 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 71.5 | 1.1e-22 | 162 | 263 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrse 89
f+k+lt sd++++g +++ +++a+e+ + + + +++l+ +d++ ++W++++i+r++++r++l++GW+ Fv++++L +gD ++F + ++
Csa02g070050.1 162 FCKTLTASDTSTHGGFSVLRRHADEClppldmSRQ-PPTQELVAKDLHANEWRFRHIFRGQPRRHLLQSGWSVFVSSKRLVAGDAFIFL--RGEN 253
99*****************************7333.34459************************************************..4499 PP
E..EEEEE-S CS
B3 90 felvvkvfrk 99
+el+v+v+r+
Csa02g070050.1 254 GELRVGVRRA 263
99*****996 PP
| |||||||
| 2 | Auxin_resp | 121.1 | 8.1e-40 | 288 | 370 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
a+ha+st+++F+v+Y+Pr+s+seF+v+++++++++k+++s+GmRfkm+fe+e+++e+r++Gt+vg++d+dp+rWp+SkWrsLk
Csa02g070050.1 288 AWHAISTGTMFTVYYKPRTSPSEFIVPFDQYMESVKNNYSIGMRFKMRFEGEEAPEQRFTGTIVGTEDSDPTRWPKSKWRSLK 370
89*******************************************************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 1.2E-40 | 158 | 277 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 3.14E-40 | 159 | 291 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 1.46E-20 | 160 | 262 | No hit | No description |
| SMART | SM01019 | 4.1E-22 | 162 | 264 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 1.5E-20 | 162 | 263 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 11.42 | 162 | 264 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 4.5E-37 | 288 | 370 | IPR010525 | Auxin response factor |
| Pfam | PF02309 | 1.2E-9 | 710 | 818 | IPR033389 | AUX/IAA domain |
| SuperFamily | SSF54277 | 2.11E-11 | 731 | 810 | No hit | No description |
| PROSITE profile | PS51745 | 24.965 | 733 | 817 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008285 | Biological Process | negative regulation of cell proliferation | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0010227 | Biological Process | floral organ abscission | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 860 aa Download sequence Send to blast |
MASSEVSMKG NRGGDNFSSS AFTDPKETTR NVSVPGEGQI SHSTRSERAV DPEAALYREL 60 WHACAGPLVT VPRQDDRVFY FPQGHIEQVE ASTNQAAEQQ MPLYDLPSKL LCRVINVDLK 120 AEADTDEVYA QITLLPEANQ DENAIEKEAP PPPPPRFQVH SFCKTLTASD TSTHGGFSVL 180 RRHADECLPP LDMSRQPPTQ ELVAKDLHAN EWRFRHIFRG QPRRHLLQSG WSVFVSSKRL 240 VAGDAFIFLR GENGELRVGV RRAMRQQGNV PSSVISSHSM HLGVLATAWH AISTGTMFTV 300 YYKPRTSPSE FIVPFDQYME SVKNNYSIGM RFKMRFEGEE APEQRFTGTI VGTEDSDPTR 360 WPKSKWRSLK VRWDETSSIP RPDRVSPWKI EPALAPPALS PVPMPRPKRP RSNIAPSSPD 420 SSMLTREGTA KANMDPLPAS GLSRVLQGQE YSTLRTKHTE SVECDAPENS VVWQSSADDD 480 KVDVVSGSRR YGSENWMSSA RHEPTYTDLL SGFGTSIDPS HGQRIPFYDH SSSPSMPAKK 540 LSSDPEGKFD FLANQWQMMH SGLSLKLHES PNLPAASDTS FQGRGNGKFS EYPVLNGLST 600 ENTGGNWPIR PRALNYFEEV VQAQAQAQAQ AREQVTKQPM TIQEETAKSR DGNCRLFGIP 660 LNNNMNGTDS TVYQRNNLND AAGLTQIASP KVQDLSDQSK GSKSTNEHRE QGRPFQTNYP 720 HPKDAHAKTS SSRSCTKVHK QGIALGRSVD LSKFQNYEEL IAELDTLFEF NGELMAPKKD 780 WLIVYTDDEN DMMLVGDDPW QEFCCMVRKI FIYTKEEVRK MNPGTLSCRS EEEAVVGEGS 840 DAKDAKSASN PSLSSAGNS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 1e-180 | 55 | 392 | 18 | 354 | Auxin response factor 1 |
| 4ldw_A | 1e-180 | 55 | 392 | 18 | 354 | Auxin response factor 1 |
| 4ldw_B | 1e-180 | 55 | 392 | 18 | 354 | Auxin response factor 1 |
| 4ldx_A | 1e-180 | 55 | 392 | 18 | 354 | Auxin response factor 1 |
| 4ldx_B | 1e-180 | 55 | 392 | 18 | 354 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). Could act as transcriptional activator or repressor. Formation of heterodimers with Aux/IAA proteins may alter their ability to modulate early auxin response genes expression. Promotes flowering, stamen development, floral organ abscission and fruit dehiscence. Functions independently of ethylene and cytokinin response pathways. May act as a repressor of cell division and organ growth. {ECO:0000269|PubMed:12036261, ECO:0000269|PubMed:15960614, ECO:0000269|PubMed:16176952, ECO:0000269|PubMed:16339187}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00574 | DAP | Transfer from AT5G62000 | Download |
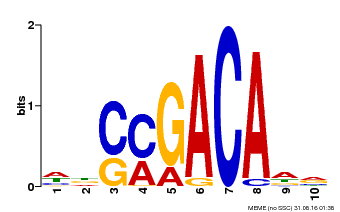
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Csa02g070050.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK221252 | 0.0 | AK221252.1 Arabidopsis thaliana mRNA for ARF1-binding protein, complete cds, clone: RAFL24-07-O11. | |||
| GenBank | AK221254 | 0.0 | AK221254.1 Arabidopsis thaliana mRNA for ARF1-binding protein, complete cds, clone: RAFL24-09-L23. | |||
| GenBank | AK221274 | 0.0 | AK221274.1 Arabidopsis thaliana mRNA for ARF1-binding protein, complete cds, clone: RAFL24-16-M02. | |||
| GenBank | AK221305 | 0.0 | AK221305.1 Arabidopsis thaliana mRNA for ARF1-binding protein, complete cds, clone: RAFL24-31-F22. | |||
| GenBank | BT000784 | 0.0 | BT000784.1 Arabidopsis thaliana clone RAFL07-10-G12 (R10939) auxin response factor (At5g62000) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010458034.1 | 0.0 | PREDICTED: auxin response factor 2 isoform X1 | ||||
| Swissprot | Q94JM3 | 0.0 | ARFB_ARATH; Auxin response factor 2 | ||||
| TrEMBL | A0A178UUC3 | 0.0 | A0A178UUC3_ARATH; Auxin response factor | ||||
| STRING | XP_010458034.1 | 0.0 | (Camelina sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5348 | 26 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62000.3 | 0.0 | auxin response factor 2 | ||||



