 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Csa02g045180.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Camelina
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 684aa MW: 75895.4 Da PI: 6.5117 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 57.5 | 2.3e-18 | 61 | 115 | 2 | 56 |
T--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 2 rkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+ +++t++q++eLe++F+ +++p++++r eL kkl L+ +q+k+WFqNrR+++k
Csa02g045180.1 61 TRYHRHTSYQIQELESFFKVCPHPNEKQRLELGKKLTLESKQIKFWFQNRRTQMK 115
334567999*******************************************999 PP
| |||||||
| 2 | START | 158.6 | 5e-50 | 228 | 428 | 4 | 206 |
HHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEEEEECTT...... CS
START 4 eeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetla....kaetlevissg...... 88
ea++el+k+a+++ p+W++ s e++ +v+ r++g+v +++ lv +l++++ +W e ++ a+t+evis+g
Csa02g045180.1 228 MEAMDELLKLAELDNPLWSSKS--EKEAIVRPAC----------TREIGLVLINSVALVDSLMETN-KWAEIFEcivaVASTVEVISNGsdgsrn 309
689******************9..7777777777..........69********************.**************************** PP
EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHHHHH CS
START 89 galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppesssvvRaellpSgiliepksnghskvtwvehvdlkgrlphwllr 182
g lqlm+ae+q++splvp R f+Ry++q+g+g w++vdvS d +++++ +s+ ++++pSg++i++ +ng skvtw+eh +++++++h+l++
Csa02g045180.1 310 GSLQLMQAEFQVMSPLVPiRQKKFLRYCKQHGDGLWAVVDVSYDINRENEHLKSYGGSKRFPSGCIIQDIGNGCSKVTWIEHLEYEESHIHSLYQ 404
**************************************************9******************************************** PP
HHHHHHHHHHHHHHHHHTXXXXXX CS
START 183 slvksglaegaktwvatlqrqcek 206
+l +s++ ga +w+atlqrqce+
Csa02g045180.1 405 PLFGSSVGLGATKWLATLQRQCES 428
**********************95 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.09E-18 | 41 | 117 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.1E-19 | 44 | 117 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 3.4E-15 | 56 | 121 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.26E-16 | 57 | 117 | No hit | No description |
| PROSITE profile | PS50071 | 16.811 | 57 | 117 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 4.1E-16 | 61 | 115 | IPR001356 | Homeobox domain |
| PROSITE profile | PS50848 | 39.134 | 216 | 431 | IPR002913 | START domain |
| CDD | cd08875 | 3.65E-99 | 222 | 427 | No hit | No description |
| SuperFamily | SSF55961 | 5.05E-28 | 223 | 429 | No hit | No description |
| SMART | SM00234 | 2.6E-37 | 225 | 428 | IPR002913 | START domain |
| Pfam | PF01852 | 2.5E-44 | 228 | 428 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 5.6E-18 | 453 | 675 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 684 aa Download sequence Send to blast |
MNGDLDVDMS RGDFNPSFYL GRLKDDEFES RSLCDDDSFD AMSGDENKQE QRPNKKKKKR 60 TRYHRHTSYQ IQELESFFKV CPHPNEKQRL ELGKKLTLES KQIKFWFQNR RTQMKTQLER 120 HENVILRQEN EKLRLENSFL KESMRGSLCI DCGGAVIPGE VSFEQHQLRI ENAKLKDELD 180 RICALANRFI GGSISLEQPS NGGNGSQHLP IGNGFSGGTS LMLMDLTMEA MDELLKLAEL 240 DNPLWSSKSE KEAIVRPACT REIGLVLINS VALVDSLMET NKWAEIFECI VAVASTVEVI 300 SNGSDGSRNG SLQLMQAEFQ VMSPLVPIRQ KKFLRYCKQH GDGLWAVVDV SYDINRENEH 360 LKSYGGSKRF PSGCIIQDIG NGCSKVTWIE HLEYEESHIH SLYQPLFGSS VGLGATKWLA 420 TLQRQCESFT NLLSSQDHTG LSLAGTKSIL KLAQRMKVNF YSGITASSVH KWEKLNAENV 480 GQDTRILTRK SLEPSGIVLS AATSLWLPVT QQRLFEFLCD GKCRNQWDIL SNGASMETTL 540 LVPKGQREGS CVSLLRAAGK DQNESSMLIL QETWNDASGA LVVYAPVDIP SMNVVMSGGD 600 SAYVALLPSG FSILPDGSSL SDQINTNGGL VNQESKGCLL TVGFQILVNS LPSAKLNVES 660 VETVNNLIAC TIHKIRAALR IPA* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that binds to the DNA sequence 5'-GCATTAAATGC-3'. {ECO:0000269|PubMed:16778018}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00559 | DAP | Transfer from AT5G52170 | Download |
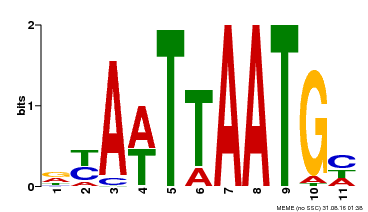
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Csa02g045180.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB025603 | 1e-166 | AB025603.1 Arabidopsis thaliana genomic DNA, chromosome 5, BAC clone:F17P19. | |||
| GenBank | CP002688 | 1e-166 | CP002688.1 Arabidopsis thaliana chromosome 5 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_019091179.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein HDG7-like isoform X2 | ||||
| Swissprot | Q9LTK3 | 0.0 | HDG7_ARATH; Homeobox-leucine zipper protein HDG7 | ||||
| TrEMBL | R0F0C9 | 0.0 | R0F0C9_9BRAS; Uncharacterized protein | ||||
| STRING | XP_010444530.1 | 0.0 | (Camelina sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1128 | 27 | 105 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G52170.1 | 0.0 | homeodomain GLABROUS 7 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



