 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Csa00430s030.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Camelina
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 828aa MW: 95093.7 Da PI: 6.8386 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 89.2 | 5.7e-28 | 68 | 155 | 2 | 91 |
FAR1 2 fYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkktekerrtraetrtgCkaklkvkkekdgkwevtkleleHnHelap 91
fY+eYAk++GF++++++s++sk+++e+++++f Cs++g + e++ + +++r+++ ++t+Cka+++vk++ dgkw ++++ ++HnHel p
Csa00430s030.1 68 FYQEYAKSMGFTTSIKNSRRSKKTKEFIDAKFACSRYGVTPETEGS--SSSRRSSIKKTDCKASMHVKRRPDGKWIIHEFIKDHNHELLP 155
9***************************************999888..8999999********************************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 8.3E-26 | 68 | 155 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 4.2E-25 | 276 | 368 | IPR018289 | MULE transposase domain |
| Pfam | PF04434 | 8.6E-8 | 556 | 591 | IPR007527 | Zinc finger, SWIM-type |
| PROSITE profile | PS50966 | 9.038 | 556 | 592 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 2.3E-7 | 567 | 594 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010018 | Biological Process | far-red light signaling pathway | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 828 aa Download sequence Send to blast |
MDLQENQVSD HAGDECMVDI MVDSSSSRDV GIADDLKAGG DLGFSGDVQD HEPRNGINFE 60 SNDAAYLFYQ EYAKSMGFTT SIKNSRRSKK TKEFIDAKFA CSRYGVTPET EGSSSSRRSS 120 IKKTDCKASM HVKRRPDGKW IIHEFIKDHN HELLPALAYH FRIQRNVKLA EKNNIDILHA 180 VSERTRKMYV EMSRQSGGYK NNGPLLQSDV NSQVGKGRYL ALEEGDSQVL LEYFKRIKKE 240 NPKFFYAIDL NEEQRLRNLF WADAKSRDDY LSFNDVVSFD TTYVKFNDKL PLALFIGVNH 300 HSQPMLLGCA LVADESKETF VWLIKTWLRA MGGRAPSAIL TDQDKFLMAA VSELLPNTRH 360 CIALWHVIEK IPEYFSIVMK RHENFLKKFN ECIFRSWTDD EFDMRWWKMV SQFGLENDEW 420 LLWLHEHRQK WVPTFMSGVF LAGMSTSQRS ESVNAFFDKY VHKKITLKEF LRQYGAILQN 480 RYEEESVADF DTCHKQPALK SPSPWEKQMA TTYTHTIFKK FQVEVLGVVA CHPRKEKEDD 540 NMATFRVQDV EKDDEFLVTW SKSKSELCCF CRMFEYKGYL CRHALMILQM CGFASIPPQY 600 ILKRWTKDAK SGVLAGEGAD QIQTRVQRYN DLCSRAIELS EEGCVSEENY NIVLRTLVET 660 LKNCVDLNNA RNSMIESSSQ LNNGAHEEES QEMSGVKATK KKTVCRKRKG QQEADQMLES 720 QQSLQPMETI SSEGMGMNGY YGPQQNGQGL LNLMEPPHEG YYVNQRTIQG LGQLNSIAPV 780 QDSFFANQQA MPGLGQMDFR PPPNFNYNLQ EEHLRSAQLP GNSSRQL* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
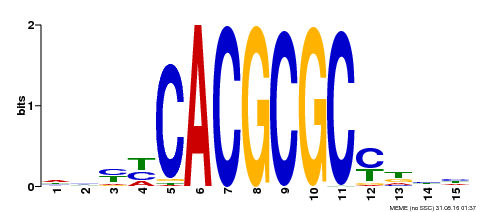
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00434 | DAP | Transfer from AT4G15090 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Csa00430s030.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. {ECO:0000269|PubMed:18033885}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF159587 | 0.0 | AF159587.1 Arabidopsis thaliana far-red impaired response protein (FAR1) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010495886.1 | 0.0 | PREDICTED: protein FAR-RED IMPAIRED RESPONSE 1-like | ||||
| Refseq | XP_010495887.1 | 0.0 | PREDICTED: protein FAR-RED IMPAIRED RESPONSE 1-like | ||||
| Refseq | XP_010495888.1 | 0.0 | PREDICTED: protein FAR-RED IMPAIRED RESPONSE 1-like | ||||
| Refseq | XP_010495889.1 | 0.0 | PREDICTED: protein FAR-RED IMPAIRED RESPONSE 1-like | ||||
| Swissprot | Q9SWG3 | 0.0 | FAR1_ARATH; Protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| TrEMBL | A0A178UYQ0 | 0.0 | A0A178UYQ0_ARATH; FAR1 | ||||
| STRING | XP_010495886.1 | 0.0 | (Camelina sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM118 | 26 | 253 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G15090.1 | 0.0 | FAR1 family protein | ||||



