 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_012575292.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Cicereae; Cicer
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 550aa MW: 60970.6 Da PI: 4.6006 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 50.8 | 3.8e-16 | 39 | 86 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT Ed++lv++vk+ G g+W+++ + g+ R++k+c++rw ++l
XP_012575292.1 39 KGPWTSAEDDILVNYVKKNGEGNWNAVQKQTGLLRCGKSCRLRWANHL 86
79******************************99**********9986 PP
| |||||||
| 2 | Myb_DNA-binding | 46.6 | 7.7e-15 | 92 | 135 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
+g++ +eE++l+++++ +lG++ W++ a+ ++ gRt++++k++w++
XP_012575292.1 92 KGAFNAEEELLIAELHSKLGNR-WARMAAQLP-GRTDNEIKNYWNT 135
799*******************.*********.***********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 14.682 | 34 | 86 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 8.59E-28 | 38 | 133 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 4.7E-13 | 38 | 88 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.1E-14 | 39 | 86 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.2E-21 | 40 | 92 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 4.80E-10 | 41 | 86 | No hit | No description |
| PROSITE profile | PS51294 | 21.725 | 87 | 141 | IPR017930 | Myb domain |
| SMART | SM00717 | 1.3E-14 | 91 | 139 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 8.8E-14 | 92 | 135 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.3E-22 | 93 | 139 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 4.67E-10 | 94 | 135 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009789 | Biological Process | positive regulation of abscisic acid-activated signaling pathway | ||||
| GO:0043068 | Biological Process | positive regulation of programmed cell death | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0045926 | Biological Process | negative regulation of growth | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 550 aa Download sequence Send to blast |
MMRRMKKEIE DGMLLNDHTE PQLNDEGYEG SVCGVVLKKG PWTSAEDDIL VNYVKKNGEG 60 NWNAVQKQTG LLRCGKSCRL RWANHLRPNL KKGAFNAEEE LLIAELHSKL GNRWARMAAQ 120 LPGRTDNEIK NYWNTRVKRC QRAGLPVYPP GVQQETPRDQ STGGINGDHK LHPDFLQKKS 180 YDDIHDAIFD SLKDNQGILP YVPELCDISA YGNMLNGFDS YQYCSFLPAT SNHKILLEST 240 MPFLDSSGNN NRSGFYPFDH IQDNTSDNLT QSFGMQSPLD PGLSSHSSMC YSHSLKNGNS 300 STSKPFEAVK LELPSLQYSE IDLGSWGTSS TPPLLESIED FIQSDTPIST LESDCSSPQN 360 SGLLDALLYP AKTLGSSKNN FYDKCSNSSI ATPGDRADSS ALNMYEPEWE EYADPVPPFG 420 ASSILNEFPA VSANRNSLDE QPPVQNFDGN IVKSEYADQI WTPDSENQIM SLLINSRPDF 480 LLASDWYEPA SGQLTNQSIA TDATSPFLGD HLATDNKHMT AGTSISNHVW EFGSCAWNNM 540 PAVCHMSDLG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 8e-29 | 37 | 139 | 5 | 106 | B-MYB |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00490 | DAP | Transfer from AT5G06100 | Download |
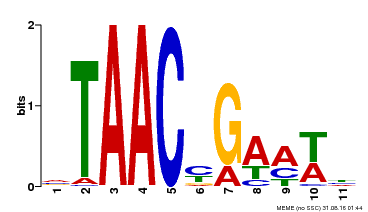
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_012575292.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004514307.1 | 0.0 | transcription factor MYB33-like | ||||
| Refseq | XP_012575292.1 | 0.0 | transcription factor MYB33-like | ||||
| TrEMBL | A0A1S2Z3B5 | 0.0 | A0A1S2Z3B5_CICAR; transcription factor MYB33-like | ||||
| STRING | XP_004514304.1 | 0.0 | (Cicer arietinum) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06100.3 | 1e-112 | myb domain protein 33 | ||||



