 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_004503803.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Cicereae; Cicer
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 867aa MW: 96842.6 Da PI: 6.5142 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 74.3 | 1.4e-23 | 166 | 267 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrse 89
f+k+lt sd++++g +++ +++a+e+ + k++ +++l+ +d++g+ W++++i+r++++r++l++GW+ Fv++++L +gD ++F + ++
XP_004503803.1 166 FCKTLTASDTSTHGGFSVLRRHADEClppldmS-KQPPTQELVAKDLHGNDWRFRHIFRGQPRRHLLQSGWSVFVSSKRLVAGDAFIFL--RGEN 257
99*****************************63.4445569************************************************..4499 PP
E..EEEEE-S CS
B3 90 felvvkvfrk 99
+el+v+v+r+
XP_004503803.1 258 GELRVGVRRA 267
99*****996 PP
| |||||||
| 2 | Auxin_resp | 118.2 | 6.6e-39 | 292 | 374 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
a+ha+ t+++F+v+Y+Pr+s++eF+v++++++++lk+++++GmRfkm+fe+e+++e+r++Gt+vg++d+d+ rWp+SkWr+Lk
XP_004503803.1 292 AWHAVLTGTMFTVYYKPRTSPAEFIVPYDQYMESLKNNYTIGMRFKMRFEGEEAPEQRFTGTIVGIEDSDSKRWPKSKWRCLK 374
79*******************************************************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 4.1E-41 | 162 | 281 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 1.22E-40 | 162 | 295 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 7.97E-21 | 164 | 266 | No hit | No description |
| PROSITE profile | PS50863 | 11.885 | 166 | 268 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 4.1E-22 | 166 | 268 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 1.7E-21 | 166 | 267 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 1.6E-36 | 292 | 374 | IPR010525 | Auxin response factor |
| Pfam | PF02309 | 1.3E-9 | 728 | 832 | IPR033389 | AUX/IAA domain |
| SuperFamily | SSF54277 | 1.86E-10 | 742 | 821 | No hit | No description |
| PROSITE profile | PS51745 | 23.773 | 744 | 826 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008285 | Biological Process | negative regulation of cell proliferation | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0010227 | Biological Process | floral organ abscission | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 867 aa Download sequence Send to blast |
MFGMRDLEEE RERRENMATS EVTMKGNCVN GKGETNDAVG DGQQNGSSST SSGREAEAAL 60 YRELWHACAG PLVTVPREGE LVFYFPQGHI EQVEASTNQV AEQHMPVYDL RPKILCRVIN 120 VMLKAEPDTD EVFAQVTLVP EPNQDENAVE KEPPPAPPPR FHVHSFCKTL TASDTSTHGG 180 FSVLRRHADE CLPPLDMSKQ PPTQELVAKD LHGNDWRFRH IFRGQPRRHL LQSGWSVFVS 240 SKRLVAGDAF IFLRGENGEL RVGVRRAMRQ QGNVPSSVIS SHSMHLGVLA TAWHAVLTGT 300 MFTVYYKPRT SPAEFIVPYD QYMESLKNNY TIGMRFKMRF EGEEAPEQRF TGTIVGIEDS 360 DSKRWPKSKW RCLKVRWDET SNIPRPERVS PWKIEPALAP PALNPLPMPR PKRPRSNVVP 420 SSPDSSVLTR EASSKVSMDP LPASGFPRVL QGQESSTLRG NLADSHESYA EKSVVWPPAA 480 DDEKIDAVST SRRYGSENWM SVSRQEPTYS DLLSGFGAVR GDNSSQPLFV DQSGHVANLG 540 RKSMLDRDGK HNVFSQWPVM PPGLSLNFLH SNTKGSPQGI PGNMRYSAFG DYSVLQGHKV 600 ESPHGNFLMP PPPPTQYDSP HLRELSHDSP HLRELSQKQI SAKTGEVAKP KDSVCKLFGF 660 SLLSSLNTPD PSIPQRNVTS ESVSHMHLAS QQHQTFENDQ KCEHSKSSKP ADNVVVVDDQ 720 GKLLQTSQPQ VKDVQLKPYN ASARSCTKVH KKGIALGRSV DLTKFTDYDE LIAELDQLFE 780 FGGELISPQK EWLVVYTDNE DDMMLVGDDP WQEFCSMVRK IYIYPKEEIQ KMSPGTLSSK 840 NEENHSASEG TDAQETKCQP NQSTSDA |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 1e-175 | 46 | 396 | 2 | 354 | Auxin response factor 1 |
| 4ldw_A | 1e-175 | 46 | 396 | 2 | 354 | Auxin response factor 1 |
| 4ldw_B | 1e-175 | 46 | 396 | 2 | 354 | Auxin response factor 1 |
| 4ldx_A | 1e-175 | 46 | 396 | 2 | 354 | Auxin response factor 1 |
| 4ldx_B | 1e-175 | 46 | 396 | 2 | 354 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). Could act as transcriptional activator or repressor. Formation of heterodimers with Aux/IAA proteins may alter their ability to modulate early auxin response genes expression. Promotes flowering, stamen development, floral organ abscission and fruit dehiscence. Functions independently of ethylene and cytokinin response pathways. May act as a repressor of cell division and organ growth. {ECO:0000269|PubMed:12036261, ECO:0000269|PubMed:15960614, ECO:0000269|PubMed:16176952, ECO:0000269|PubMed:16339187}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00574 | DAP | Transfer from AT5G62000 | Download |
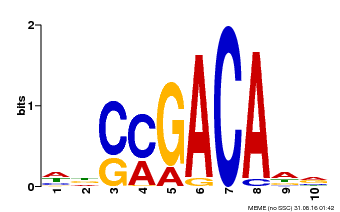
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_004503803.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004503803.1 | 0.0 | auxin response factor 2B | ||||
| Swissprot | Q94JM3 | 0.0 | ARFB_ARATH; Auxin response factor 2 | ||||
| TrEMBL | A0A1S2YG19 | 0.0 | A0A1S2YG19_CICAR; Auxin response factor | ||||
| STRING | XP_004503803.1 | 0.0 | (Cicer arietinum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3281 | 33 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62000.3 | 0.0 | auxin response factor 2 | ||||



