 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_004499875.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Cicereae; Cicer
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 342aa MW: 39257.1 Da PI: 5.8289 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 50.3 | 5.6e-16 | 32 | 79 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g W++eEde+lv++ G g+W+ Iar g+ R++k+c++rw +yl
XP_004499875.1 32 KGLWSPEEDEKLVRYMITKGQGCWSDIARNAGLQRCGKSCRLRWINYL 79
678*******************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 52.4 | 1.2e-16 | 85 | 129 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
rg+++++E+el+++++ lG++ W+ Ia++++ gRt++++k++w++
XP_004499875.1 85 RGAFSPQEEELIIHLHSILGNR-WSQIAARLP-GRTDNEIKNFWNST 129
89********************.*********.***********986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 17.36 | 27 | 79 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.27E-28 | 30 | 126 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 9.1E-13 | 31 | 81 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.9E-14 | 32 | 79 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.0E-23 | 33 | 86 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 7.61E-11 | 35 | 79 | No hit | No description |
| PROSITE profile | PS51294 | 26.126 | 80 | 134 | IPR017930 | Myb domain |
| SMART | SM00717 | 1.8E-16 | 84 | 132 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.9E-15 | 85 | 129 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.6E-26 | 87 | 135 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.57E-11 | 87 | 130 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:2000652 | Biological Process | regulation of secondary cell wall biogenesis | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 342 aa Download sequence Send to blast |
MQMRKPDLIT GINNKMNNNN NNNNISKNKL RKGLWSPEED EKLVRYMITK GQGCWSDIAR 60 NAGLQRCGKS CRLRWINYLR PDLKRGAFSP QEEELIIHLH SILGNRWSQI AARLPGRTDN 120 EIKNFWNSTL KKRLKTNNNN TSQSPSDSSD QQIDVFDMKN NMIMPINEQD LMTLCMDNSS 180 SSTSSSSIQS MQMHAMVLSD QFDHPFSLLS NINNIPTCLT QDGNMVVDDI GLHSDDHYGI 240 LEANNKMGLL ENDFELPPLE SRISMEDKSM IPLINHNVMK SYNINHFNNS CFNNTNQIQN 300 SKVEDFFGFG NHGQGENLRM GEWDFEGLMQ DMSYFPTLDF QV |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 5e-28 | 30 | 136 | 5 | 110 | B-MYB |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 129 | 135 | LKKRLKT |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
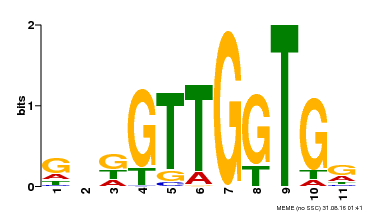
| MP00336 | DAP | Transfer from AT3G08500 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_004499875.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004499875.1 | 0.0 | transcription factor MYB83-like | ||||
| TrEMBL | A0A1S2Y5Z6 | 0.0 | A0A1S2Y5Z6_CICAR; transcription factor MYB83-like | ||||
| STRING | XP_004499875.1 | 0.0 | (Cicer arietinum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF774 | 34 | 128 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G08500.1 | 7e-69 | myb domain protein 83 | ||||



