 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_004499538.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Cicereae; Cicer
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 535aa MW: 59806.9 Da PI: 7.3209 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 54.9 | 1.5e-17 | 342 | 388 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
hn ErrRRdriN+++ L++l+P++ K +Ka++Le+A+eY+ksLq
XP_004499538.1 342 VHNLSERRRRDRINEKMKALQQLIPHS-----SKTDKASMLEEAIEYLKSLQ 388
6*************************8.....6******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.280.10 | 1.5E-19 | 335 | 396 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 3.14E-20 | 336 | 404 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 18.47 | 338 | 387 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 8.99E-18 | 341 | 392 | No hit | No description |
| Pfam | PF00010 | 4.8E-15 | 342 | 388 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.3E-18 | 344 | 393 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009704 | Biological Process | de-etiolation | ||||
| GO:0010161 | Biological Process | red light signaling pathway | ||||
| GO:0010244 | Biological Process | response to low fluence blue light stimulus by blue low-fluence system | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0010928 | Biological Process | regulation of auxin mediated signaling pathway | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 535 aa Download sequence Send to blast |
MNDSVSEWNF ESDTYVTNQK KSIGLDHELV ELLWQNGQVV LHSQTNRKPI TSRHVQKNLQ 60 STTLRTNETF GSSSNLIQDD ETVSWIHYPL EDPLEQELCS NLLSELSPCD DVVESYKQIR 120 NFEEGKFSKL DASSDPHANI NSQPSNLKPS FVQEFSKPDQ RFHHAPDSSH KILNFSHFSR 180 PPNVSSVTGK GDAHFRDKVT SGNLSQGDIR ECSAMTVVSS YCGSNHIPQD QDVSRVSSNG 240 VWNTTLSAEP KPVKDDIHKT IPRHEKGKSE MIEPTVTSSS GGSGSSLGKT SSLSTRSHGQ 300 KRKIIDVEDS VEQSEDTELK SAVRNKVSQR SGSARRNRAA EVHNLSERRR RDRINEKMKA 360 LQQLIPHSSK TDKASMLEEA IEYLKSLQLQ LQVMWMGSGM TPMMLPGFQH YMSQMGMGMT 420 APSFPPLQNP MQLQRIPLDQ SVQTPNQTLM CPNPILGAFN YQNQIQNPAL SDQYARYMGY 480 HLMQNASQPM SMYRYGPQGV QQTQAMIPPS NSSGPIRGVV NVDDSVSGKM GRPLT |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 346 | 351 | ERRRRD |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00606 | ChIP-seq | Transfer from AT2G43010 | Download |
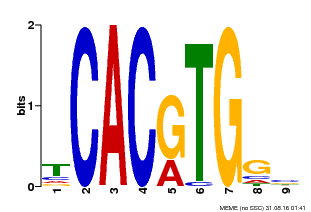
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_004499538.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015038 | 9e-54 | AP015038.1 Vigna angularis var. angularis DNA, chromosome 5, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004499538.1 | 0.0 | transcription factor PIF4-like isoform X1 | ||||
| TrEMBL | A0A1S2Y588 | 0.0 | A0A1S2Y588_CICAR; transcription factor PIF4-like isoform X1 | ||||
| STRING | XP_004499538.1 | 0.0 | (Cicer arietinum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF8146 | 29 | 44 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G43010.2 | 2e-62 | phytochrome interacting factor 4 | ||||



