 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_004498462.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Cicereae; Cicer
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 347aa MW: 38518.7 Da PI: 6.7831 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 56.1 | 4.9e-18 | 239 | 273 | 1 | 35 |
GATA 1 CsnCgttkTplWRrgpdgnktLCnaCGlyyrkkgl 35
C +C t kTp+WR gp g+ktLCnaCG++y++ +l
XP_004498462.1 239 CLHCATDKTPQWRTGPKGPKTLCNACGVRYKSGRL 273
99*****************************9885 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF016992 | 1.8E-79 | 14 | 329 | IPR016679 | Transcription factor, GATA, plant |
| SMART | SM00401 | 6.6E-17 | 233 | 283 | IPR000679 | Zinc finger, GATA-type |
| PROSITE profile | PS50114 | 12.347 | 233 | 269 | IPR000679 | Zinc finger, GATA-type |
| SuperFamily | SSF57716 | 2.14E-15 | 234 | 297 | No hit | No description |
| Gene3D | G3DSA:3.30.50.10 | 1.5E-14 | 235 | 270 | IPR013088 | Zinc finger, NHR/GATA-type |
| CDD | cd00202 | 1.75E-13 | 238 | 285 | No hit | No description |
| Pfam | PF00320 | 6.1E-16 | 239 | 273 | IPR000679 | Zinc finger, GATA-type |
| PROSITE pattern | PS00344 | 0 | 239 | 264 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009416 | Biological Process | response to light stimulus | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 347 aa Download sequence Send to blast |
MEGTSNFFQT TFCQQQLSSD NNQTNNTTST TTTADQFIVE DLLNLSNDDV IIAETNLDAA 60 IGTSTNSSTI INSCNSSFSG CDHNLVHDIG GHNLSDCHFS GDLCVPNDDL AELEWLSNFV 120 EESFSSEDLQ KLQLISGKKV GSKDEEVHEF SQPNRNNPIF NKEVSVPAKA RSKRTRGPPC 180 DWSSRLLVLS ETASSSSESE LLIPTPTLPS TTVLPKKQKT ASRRKDNGDG GSGGDGRRCL 240 HCATDKTPQW RTGPKGPKTL CNACGVRYKS GRLVPEYRPA ASPTFVLTKH SNSHRKVLEL 300 RRQKEMVRAH QHHQHQFLHH QNMIFDISNS DEYLIHQPVG PDFRQLI |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that specifically binds 5'-GATA-3' or 5'-GAT-3' motifs within gene promoters. May be involved in the regulation of some light-responsive genes (By similarity). Transcription activator involved in xylem formation. Functions upstream of NAC030/VND7, a master switch of xylem vessel differentiation (PubMed:25265867). {ECO:0000250|UniProtKB:Q8LAU9, ECO:0000269|PubMed:25265867}. | |||||
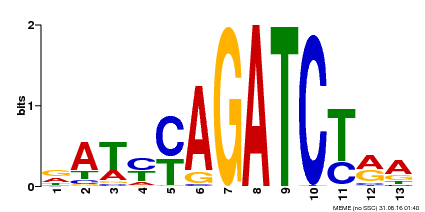
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00530 | DAP | Transfer from AT5G25830 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_004498462.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004498462.1 | 0.0 | GATA transcription factor 12-like | ||||
| Swissprot | P69781 | 8e-94 | GAT12_ARATH; GATA transcription factor 12 | ||||
| TrEMBL | A0A1S2Y4M0 | 0.0 | A0A1S2Y4M0_CICAR; GATA transcription factor | ||||
| STRING | XP_004498462.1 | 0.0 | (Cicer arietinum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1364 | 33 | 95 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G32890.1 | 9e-68 | GATA transcription factor 9 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



