 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_004495934.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Cicereae; Cicer
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 302aa MW: 34681.4 Da PI: 4.9413 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 60.1 | 5.3e-19 | 79 | 127 | 3 | 55 |
AP2 3 ykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
y+G+r+++ +g+W+AeIrdp++ r++lg+f taeeAa+a+++ ++k++g
XP_004495934.1 79 YRGIRQRP-WGKWAAEIRDPRK---GVRVWLGTFNTAEEAARAYDREARKIRG 127
9*******.**********954...4************************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:3.30.730.10 | 3.4E-32 | 78 | 136 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 23.775 | 78 | 135 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 2.42E-22 | 78 | 136 | IPR016177 | DNA-binding domain |
| SMART | SM00380 | 1.4E-38 | 78 | 141 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 6.1E-12 | 79 | 127 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 7.7E-12 | 79 | 90 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 3.41E-33 | 79 | 137 | No hit | No description |
| PRINTS | PR00367 | 7.7E-12 | 101 | 117 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008219 | Biological Process | cell death | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009735 | Biological Process | response to cytokinin | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010286 | Biological Process | heat acclimation | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0051707 | Biological Process | response to other organism | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005737 | Cellular Component | cytoplasm | ||||
| GO:0005886 | Cellular Component | plasma membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 302 aa Download sequence Send to blast |
MCGGAIIADF IPRRGGRRLT ASELWPSSFG QQKDSDFDFS QTGGDQQTPP PPPTLKRSYT 60 PTVSEQVEKP VKKQRKNLYR GIRQRPWGKW AAEIRDPRKG VRVWLGTFNT AEEAARAYDR 120 EARKIRGKKA KVNFPNDDDD YSSSIKVRRY NNPPIKPPII PLYQQQQYPK NLNLEFGYDL 180 NQTDTVHSQV GAINTDSFII SGNDETSEVG NGYSTMEFNQ NENGYFVGVV NENEKEKEKE 240 KEREIEQQEQ EAAKSEVQEL SDELMAYEDY MKFYQIPYYG GQFMPSPNIV QESVVGDLWS 300 FD |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1gcc_A | 4e-22 | 79 | 135 | 3 | 60 | ETHYLENE RESPONSIVE ELEMENT BINDING FACTOR 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00109 | PBM | Transfer from AT3G16770 | Download |
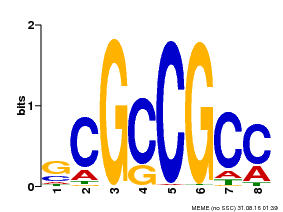
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_004495934.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB378638 | 4e-73 | AB378638.1 Lotus japonicus LjERF19 mRNA for transcription factor AP2-EREBP, complete cds. | |||
| GenBank | KJ004728 | 4e-73 | KJ004728.1 Lotus corniculatus cultivar Leo AP2/ERF (ERF054) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004495934.1 | 0.0 | ethylene-responsive transcription factor ERF113-like | ||||
| TrEMBL | A0A1S2XXC7 | 0.0 | A0A1S2XXC7_CICAR; ethylene-responsive transcription factor ERF113-like | ||||
| STRING | XP_004495934.1 | 0.0 | (Cicer arietinum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF6771 | 34 | 49 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G16770.1 | 4e-35 | ethylene-responsive element binding protein | ||||



