 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_004495851.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Cicereae; Cicer
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 428aa MW: 48410 Da PI: 6.6652 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 54.5 | 2.8e-17 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g W++eEde+l++ ++++G g+W+++++ g+ R++k+c++rw +yl
XP_004495851.1 14 KGLWSPEEDEKLLRHITKYGHGCWSSVPKQAGLQRCGKSCRLRWINYL 61
678*******************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 44.8 | 2.9e-14 | 67 | 110 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
rg ++++E+ l++++++ lG++ W+ Ia+ ++ gRt++++k+ w++
XP_004495851.1 67 RGTFSQDEENLIIELHAVLGNR-WSQIAAQLP-GRTDNEIKNLWNS 110
899*******************.*********.***********97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 4.3E-28 | 6 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 18.926 | 9 | 61 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 3.67E-29 | 12 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.7E-13 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.2E-15 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.06E-11 | 17 | 61 | No hit | No description |
| PROSITE profile | PS51294 | 25.357 | 62 | 116 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.3E-25 | 65 | 116 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 7.3E-13 | 66 | 114 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.5E-13 | 67 | 111 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.07E-8 | 69 | 109 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 428 aa Download sequence Send to blast |
MGRHSCCYKQ KLRKGLWSPE EDEKLLRHIT KYGHGCWSSV PKQAGLQRCG KSCRLRWINY 60 LRPDLKRGTF SQDEENLIIE LHAVLGNRWS QIAAQLPGRT DNEIKNLWNS CLKKKLRQKG 120 IDPVTHKPLS EVENLEDNVK SQEKVAQVSN ELNLLKSESS KSDAASCDHR TSTSLSPKAY 180 AQEMDGSGSY TKSESSNLMN SKDMFLDKFM NHQTDLMANF PLQMSFSNTE NDCLTNDSNS 240 SHWFNQTGKT FDMNSEFPFN SNSILTPTTT TSTTTMFLPN SFCCNDSSLA TIHSDNVSTP 300 CGSNYWEVNR SDNRSNNSST LNLQNNIFSS WGLADCSINS TKEAQIHDMI EHNHAEEEAK 360 WENNYLHNPI LMLASESLCN EIKPSMMQLV PHDTLGAILP HTKQQEQSQA SNIFSKDIQK 420 LRAAFGDM |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 5e-28 | 12 | 116 | 5 | 108 | B-MYB |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 111 | 117 | LKKKLRQ |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00427 | DAP | Transfer from AT4G01680 | Download |
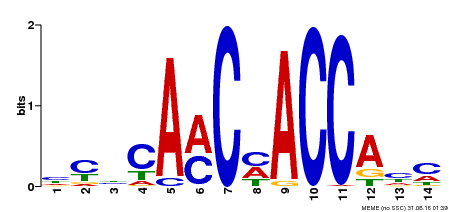
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_004495851.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT052849 | 1e-114 | BT052849.1 Medicago truncatula clone MTYFL_FM_FN_FO1G-P-4 unknown mRNA. | |||
| GenBank | BT147477 | 1e-114 | BT147477.1 Medicago truncatula clone JCVI-FLMt-12D24 unknown mRNA. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004495851.1 | 0.0 | transcription factor MYB61 | ||||
| TrEMBL | A0A1S2XYD6 | 0.0 | A0A1S2XYD6_CICAR; transcription factor MYB61 | ||||
| STRING | XP_004495851.1 | 0.0 | (Cicer arietinum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1083 | 33 | 103 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G01680.3 | 1e-93 | myb domain protein 55 | ||||



