 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_004494461.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Cicereae; Cicer
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 995aa MW: 110775 Da PI: 6.2396 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 125.6 | 2e-39 | 138 | 214 | 2 | 78 |
-SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 2 CqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
Cqve+C+adl++ak+yhrrhkvCe+hska ++lv +++qrfCqqCsrfh+l+efDe+krsCrrrLa+hn+rrrk+++
XP_004494461.1 138 CQVEDCRADLNNAKDYHRRHKVCEIHSKACKALVGNTMQRFCQQCSRFHMLQEFDEGKRSCRRRLAGHNKRRRKTNQ 214
**************************************************************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 2.7E-31 | 131 | 199 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 31.71 | 135 | 212 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 9.68E-37 | 136 | 216 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 9.1E-28 | 138 | 211 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF48403 | 9.12E-8 | 763 | 883 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 1.7E-7 | 769 | 884 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 6.14E-8 | 769 | 881 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 995 aa Download sequence Send to blast |
MEAFHMYGME GGSSDLRITG KRSLEWDLND WKWDGDVFVA SRRLSPVPEH RQFLPLPGGG 60 SSNSNSSSSC SEDLDLGNKE GERKRRVIVV EDELSLNKEA GSLSLKIGGG SAQIATWEGN 120 SGKKSRVAAG GTSSRAFCQV EDCRADLNNA KDYHRRHKVC EIHSKACKAL VGNTMQRFCQ 180 QCSRFHMLQE FDEGKRSCRR RLAGHNKRRR KTNQDAVPNG SPLNDDQTSS YLLISLLKIL 240 SNMQPDRTNQ TADQDLLTHL LRSLANQNGE QGAKNLSNLL REPENLLKEG SSSGKSEMIS 300 TLFTNCSQGS PTVTRQNQTV SISEIQHQVM HAHDARAADQ QTTSSAKPSV SNSPPAYSEA 360 RDSTAGQTKM NNFDLNDIYV DSDDGIEDIE RFPVSVNLGT SSLDYPWMQQ DSHQSSPPQT 420 SGNSDSASAQ SPSSSTGEAQ SLTDRIVFKL FGKEPSDFPL VLRAQILDWL SHSPTDIESY 480 IRPGCVILTI YLRQAEVVWE ELCFDLTSSL NRLLDVSDDD FWKTGWVHIR VQHQIAFIFN 540 GQVVIDTSLP FRSNNYSKIL SVSPIAVPAS KTAQFSVKGI NLTRPATRLL CALEGNYLVC 600 EDTHESMDQC SKDLDELQCV QFSCSVPVMN GRGFIEIEDQ GLSSSFFPFI VVEEDVCSEI 660 CVLEPLLESS DTDSDVERAG RIQAKNQAID FIHEMGWLLH RSQIKSRMVH LSSSADLFPL 720 DRFNWLMEFS MDHDWCAVVK KLLNLLLNGT VSTGDHASLY LALSDMGLLH RAVRRNSRQL 780 VELLLRYVPQ NISDTLGPED KALVNRENQN FLFRPDVVGP AGLTPLHIAA GKDGSEDVLD 840 ALTNDPCMVG IEAWKSARDS TGSTPEDYAR LRGHYTYIHL LQKKINKRQG GAHVVVDIPS 900 NLTRFVTSQN KDESSTTFEI GNAEVRNVQK QCKLCDHKLS CRTAVRKSFV YRPAMLSMVA 960 IAAVCVCVAL LFKSSPEVLY IFRPFRWESL EFGTS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 3e-30 | 128 | 211 | 1 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16095614}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
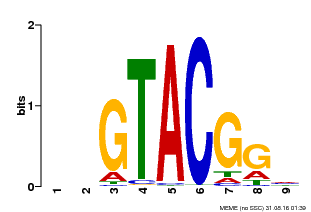
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_004494461.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004494461.1 | 0.0 | squamosa promoter-binding-like protein 1 isoform X2 | ||||
| Swissprot | Q9SMX9 | 0.0 | SPL1_ARATH; Squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | A0A1S2XV77 | 0.0 | A0A1S2XV77_CICAR; squamosa promoter-binding-like protein 1 isoform X2 | ||||
| STRING | XP_004494461.1 | 0.0 | (Cicer arietinum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1539 | 34 | 95 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



