 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_004492415.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Cicereae; Cicer
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 312aa MW: 35649.3 Da PI: 8.3519 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 24.2 | 7.6e-08 | 41 | 86 | 3 | 46 |
SS-HHHHHHHHHHHHHTTTT...-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGgg...tWktIartmgkgRtlkqcksrwqk 46
+WT+eE +l+ +a + + ++ +W ++a++++ g+t +++ ++
XP_004492415.1 41 KWTQEENKLFENALAYYDKDtpdRWIRVAEMIP-GKTVGDVIKQYRE 86
7********************************.******9999986 PP
| |||||||
| 2 | Myb_DNA-binding | 46.1 | 1.2e-14 | 148 | 192 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE+ +++ + k++G+g+W+ I+r + ++Rt+ q+ s+ qky
XP_004492415.1 148 PWTEEEHRQFLLGLKKYGKGDWRNISRNFVTTRTPTQVASHAQKY 192
8*******************************************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51293 | 8.289 | 37 | 91 | IPR017884 | SANT domain |
| SMART | SM00717 | 1.6E-7 | 38 | 90 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 5.91E-12 | 41 | 95 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 4.76E-6 | 41 | 87 | No hit | No description |
| Pfam | PF00249 | 1.6E-6 | 41 | 86 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS51294 | 20.205 | 141 | 197 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.0E-17 | 142 | 196 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 9.4E-18 | 144 | 196 | IPR006447 | Myb domain, plants |
| Gene3D | G3DSA:1.10.10.60 | 1.9E-12 | 145 | 192 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 3.5E-13 | 145 | 195 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.01E-10 | 148 | 193 | No hit | No description |
| Pfam | PF00249 | 4.2E-12 | 148 | 192 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 312 aa Download sequence Send to blast |
MFSCYMQFLS SRKMNSEIEV LSPSSYIQNS NWLFQENKGA KWTQEENKLF ENALAYYDKD 60 TPDRWIRVAE MIPGKTVGDV IKQYRELEED VCVIEAGLIP VPGYTTSSFT LDWANNQGFD 120 EFKQFCSVGG KRGASTRPTE QERKKGVPWT EEEHRQFLLG LKKYGKGDWR NISRNFVTTR 180 TPTQVASHAQ KYFIRQVTGG KDKRRSSIHD ITVVNLPETK SPLSDGSMPS PNPQKNHKLS 240 SMIKQEYDWK LPDEEAMPFV FNSTTKGNMF VAPLCGIASH ESKPPGCNVL RGIQHGCQFS 300 LPFDTILQMQ HQ |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cjj_A | 2e-16 | 39 | 107 | 8 | 76 | RADIALIS |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Involved in the dorsovental asymmetry of flowers. Promotes ventral identity. {ECO:0000269|PubMed:11937495, ECO:0000269|PubMed:9118809}. | |||||
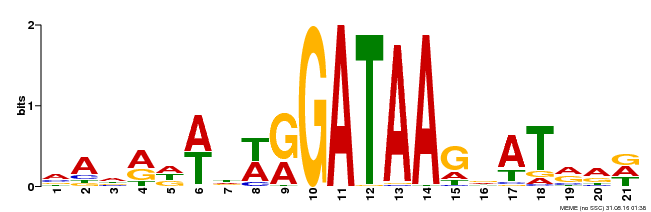
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00296 | DAP | Transfer from AT2G38090 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_004492415.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | EF468478 | 0.0 | EF468478.1 Medicago truncatula MYB transcription factor MYB33 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004492415.1 | 0.0 | transcription factor DIVARICATA-like | ||||
| Swissprot | Q8S9H7 | 1e-107 | DIV_ANTMA; Transcription factor DIVARICATA | ||||
| TrEMBL | A0A1S2XQJ5 | 0.0 | A0A1S2XQJ5_CICAR; transcription factor DIVARICATA-like | ||||
| STRING | XP_004492415.1 | 0.0 | (Cicer arietinum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF466 | 34 | 162 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G38090.1 | 1e-120 | MYB family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



