 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_004487499.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Cicereae; Cicer
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 479aa MW: 53401 Da PI: 6.5916 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 32.5 | 1.9e-10 | 174 | 215 | 4 | 45 |
XCHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 4 lkrerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaL 45
k rr+++NReAAr+sR+RKka++++Le +L++ ++L
XP_004487499.1 174 AKTLRRLAQNREAARKSRLRKKAYVQQLESSRLKLTQLEQDL 215
6889*************************9877777766666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 9.6E-7 | 171 | 237 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 9.232 | 173 | 217 | IPR004827 | Basic-leucine zipper domain |
| Gene3D | G3DSA:1.20.5.170 | 5.0E-8 | 173 | 217 | No hit | No description |
| Pfam | PF00170 | 1.2E-7 | 174 | 216 | IPR004827 | Basic-leucine zipper domain |
| CDD | cd14708 | 9.51E-22 | 175 | 225 | No hit | No description |
| SuperFamily | SSF57959 | 9.64E-7 | 175 | 217 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 178 | 193 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF14144 | 3.2E-26 | 256 | 330 | IPR025422 | Transcription factor TGA like domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 479 aa Download sequence Send to blast |
MASQRIGELD FSEPYGIHHR INNPSSSLIN DQGSGFDFGE LEEAIGLQGV KIRNDETKAS 60 AFFTGKPAAT LEMFPSWPMR FHQSSIGGTN NKSGGESSDS ALSSKNENPF EPESPLSKKA 120 SSFCSSDQLQ HLQLQQDMTN NNDVIKTASS PKSAHEKLQR KGGGSTSDNK PLDAKTLRRL 180 AQNREAARKS RLRKKAYVQQ LESSRLKLTQ LEQDLQRARS QGMFMDWSGG VGSNISSGAM 240 FDMEYARWLE EDNRLLTELR NGLEAALSDN EMRVMVEGYL CHYDEVFRVK GVTAKADVFH 300 VINGGWTSQA ERCFLWIGGF RPSEIIMMLI QQLEAVAEQQ IMGMYGLRHS SQQAEEALSQ 360 GLDQLQQSVV DTIAGAPLVD GVQQMVVAVG KLANLEGFLR QADNLRQQTL HQLCRLLTVR 420 QAARSFLVIG EYYARLRALS SLWASRSRDQ TIITSDENSC QTTTDMQMLQ PSHNHFSSF |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Together with TGA10, basic leucine-zipper transcription factor required for anther development, probably via the activation of SPL expression in anthers and via the regulation of genes with functions in early and middle tapetal development (PubMed:20805327). Required for signaling responses to pathogen-associated molecular patterns (PAMPs) such as flg22 that involves chloroplastic reactive oxygen species (ROS) production and subsequent expression of H(2)O(2)-responsive genes (PubMed:27717447). {ECO:0000269|PubMed:20805327, ECO:0000269|PubMed:27717447}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00131 | DAP | Transfer from AT1G08320 | Download |
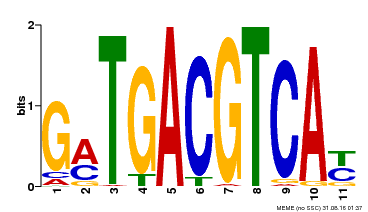
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_004487499.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by flg22 in leaves. {ECO:0000269|PubMed:27717447}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC235234 | 6e-36 | AC235234.1 Glycine max strain Williams 82 clone GM_WBb0028B23, complete sequence. | |||
| GenBank | AP015040 | 6e-36 | AP015040.1 Vigna angularis var. angularis DNA, chromosome 7, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004487499.1 | 0.0 | transcription factor TGA9-like isoform X1 | ||||
| Swissprot | Q93XM6 | 0.0 | TGA9_ARATH; Transcription factor TGA9 | ||||
| TrEMBL | A0A1S2XD58 | 0.0 | A0A1S2XD58_CICAR; transcription factor TGA9-like isoform X1 | ||||
| STRING | XP_004487499.1 | 0.0 | (Cicer arietinum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF4555 | 33 | 57 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G08320.3 | 0.0 | bZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



