 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_004485583.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Cicereae; Cicer
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1019aa MW: 114857 Da PI: 5.8073 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 179.5 | 4.1e-56 | 18 | 132 | 4 | 118 |
CG-1 4 ekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrc 98
++rwl++ ei++iL n++ +++t+e++ rp+sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+hekLKvg+v+vl+cyYah+een++fqrr+
XP_004485583.1 18 AQHRWLRPAEICEILCNYRMFHITSEPHIRPPSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHEKLKVGSVDVLHCYYAHGEENENFQRRS 112
49********************************************************************************************* PP
CG-1 99 ywlLeeelekivlvhylevk 118
yw+L+ e+ +iv+vhylevk
XP_004485583.1 113 YWMLDPEMMHIVFVHYLEVK 132
*****************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 81.847 | 11 | 137 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 7.2E-80 | 14 | 132 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 3.6E-50 | 17 | 131 | IPR005559 | CG-1 DNA-binding domain |
| SuperFamily | SSF81296 | 2.52E-17 | 434 | 519 | IPR014756 | Immunoglobulin E-set |
| Gene3D | G3DSA:2.60.40.10 | 4.9E-5 | 434 | 508 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 4.0E-7 | 434 | 510 | IPR002909 | IPT domain |
| PROSITE profile | PS50297 | 20.394 | 605 | 738 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 5.75E-20 | 610 | 728 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 9.4E-9 | 616 | 695 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 2.3E-20 | 618 | 729 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 5.07E-16 | 623 | 726 | No hit | No description |
| SMART | SM00248 | 3600 | 634 | 663 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 8.63 | 634 | 666 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 10.606 | 667 | 699 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 3.5E-4 | 667 | 696 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 1300 | 706 | 735 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 1.65E-7 | 838 | 893 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.15 | 842 | 864 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 7.712 | 844 | 872 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.004 | 846 | 863 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0024 | 865 | 887 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.432 | 866 | 890 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 5.3E-5 | 868 | 887 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1019 aa Download sequence Send to blast |
MAEPPSYGLD MQQLQFEAQH RWLRPAEICE ILCNYRMFHI TSEPHIRPPS GSLFLFDRKV 60 LRYFRKDGHN WRKKKDGKTV KEAHEKLKVG SVDVLHCYYA HGEENENFQR RSYWMLDPEM 120 MHIVFVHYLE VKGNKSNIGG NSDCSVPSLS TDPMSPTSSL ASLREDADSG DHGQSSVSGT 180 DYIPLVDMDK YRGNDATCID GLKAHDMASW DTVLQSTGEL HADPSLVSFP SIPSSSLANI 240 LDQEQNIFGD FSMSRSDLTI GAGSSQPLQS NWQIPFEDNT GHMPSLTQSL SLEFGSDYGT 300 GLLGNEAQNE SSEIDPVMFS FHGEPKEKLA QQNYLEKKVE GHLQDELKSN CANEVHIEET 360 INYPLSVRRT LLDSNESLKK VDSFSRWITK ALGEVDNLNM QSSPGISWST DECGHVIDDT 420 SLSPSLSQDQ LYSINDFSPK WAYAGSDTEV LIIGSFLKSQ PEVTTYNWSC MFGEVEVPAE 480 VVANGILCCQ APPHKVGRVP FYVTCSNRLA CSEVREFDFR EGYSSNVDYT DFFNSSNDML 540 LHLRLDKFLS LKPVHPSNQA FEGDMEKINL IFKLISLREE EDYSSKEEKT VEMNISRHKV 600 KEHQFHRQFK ENLYSWLLHK VTESGKGPNV LDKDGQGVLH LAAVLGYYWA ITPILIAGVN 660 VNFRDVNGWT ALHWAASCGR ERTVAVLVSM GADCGALTDP SPEFPSGRTA ADLASSNGHK 720 GISGFLAESS LTSHLESLTV DDKQKGGQQE ISGTKAVQTV SERTATPVVY NDMPDGLCLK 780 DSLTAVRNAT QAADRIHQVF RMQSFQRKQL TQYEDDEFGL SDQRALSLLA SKVCKSGQRD 840 GLVNVAATQI QKKFRGWKKR KEFLIIRERI VKIQAHVRGH QVRKQYKTII WSVGILEKVI 900 LRWRRKGSGL RGFRPDTLNK APSQQSDSLK EDDYDYLKEG RKQKEEKIEK ALSRVKSMVQ 960 YPEARAQYRR VLNVVEDFRQ KKDSNMGLIS SEETVDGVED LIDIDMLLDD DNFIPIAFD |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5hry_A | 9e-12 | 611 | 728 | 46 | 157 | ank3C2_1 |
| 5hry_B | 9e-12 | 611 | 728 | 46 | 157 | ank3C2_1 |
| 5hry_C | 9e-12 | 611 | 728 | 46 | 157 | ank3C2_1 |
| 5hry_D | 9e-12 | 611 | 728 | 46 | 157 | ank3C2_1 |
| 5hry_E | 9e-12 | 611 | 728 | 46 | 157 | ank3C2_1 |
| 5hry_F | 9e-12 | 611 | 728 | 46 | 157 | ank3C2_1 |
| 5hry_G | 9e-12 | 611 | 728 | 46 | 157 | ank3C2_1 |
| 5hry_H | 9e-12 | 611 | 728 | 46 | 157 | ank3C2_1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
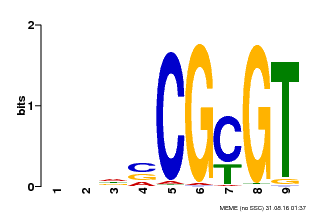
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_004485583.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC187465 | 0.0 | AC187465.3 Medicago truncatula chromosome 2 BAC clone mth2-69g14, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004485583.1 | 0.0 | calmodulin-binding transcription activator 2-like isoform X2 | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A1S2X9F5 | 0.0 | A0A1S2X9F5_CICAR; calmodulin-binding transcription activator 2-like isoform X2 | ||||
| STRING | XP_004485582.1 | 0.0 | (Cicer arietinum) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



