 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_004485582.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Cicereae; Cicer
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1023aa MW: 115281 Da PI: 5.8722 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 179.5 | 4.1e-56 | 22 | 136 | 4 | 118 |
CG-1 4 ekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrc 98
++rwl++ ei++iL n++ +++t+e++ rp+sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+hekLKvg+v+vl+cyYah+een++fqrr+
XP_004485582.1 22 AQHRWLRPAEICEILCNYRMFHITSEPHIRPPSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHEKLKVGSVDVLHCYYAHGEENENFQRRS 116
49********************************************************************************************* PP
CG-1 99 ywlLeeelekivlvhylevk 118
yw+L+ e+ +iv+vhylevk
XP_004485582.1 117 YWMLDPEMMHIVFVHYLEVK 136
*****************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 81.847 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 7.2E-80 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 3.6E-50 | 21 | 135 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 5.0E-5 | 438 | 512 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 4.0E-7 | 438 | 514 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 2.52E-17 | 438 | 523 | IPR014756 | Immunoglobulin E-set |
| PROSITE profile | PS50297 | 20.394 | 609 | 742 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 5.75E-20 | 614 | 732 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 9.4E-9 | 620 | 699 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 2.3E-20 | 622 | 733 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 5.10E-16 | 627 | 730 | No hit | No description |
| PROSITE profile | PS50088 | 8.63 | 638 | 670 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 3600 | 638 | 667 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 3.5E-4 | 671 | 700 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 10.606 | 671 | 703 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 1300 | 710 | 739 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 1.65E-7 | 842 | 897 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.15 | 846 | 868 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 7.712 | 848 | 876 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0041 | 850 | 867 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0024 | 869 | 891 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.432 | 870 | 894 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 5.3E-5 | 872 | 891 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1023 aa Download sequence Send to blast |
MAEPPSYGLG PRLDMQQLQF EAQHRWLRPA EICEILCNYR MFHITSEPHI RPPSGSLFLF 60 DRKVLRYFRK DGHNWRKKKD GKTVKEAHEK LKVGSVDVLH CYYAHGEENE NFQRRSYWML 120 DPEMMHIVFV HYLEVKGNKS NIGGNSDCSV PSLSTDPMSP TSSLASLRED ADSGDHGQSS 180 VSGTDYIPLV DMDKYRGNDA TCIDGLKAHD MASWDTVLQS TGELHADPSL VSFPSIPSSS 240 LANILDQEQN IFGDFSMSRS DLTIGAGSSQ PLQSNWQIPF EDNTGHMPSL TQSLSLEFGS 300 DYGTGLLGNE AQNESSEIDP VMFSFHGEPK EKLAQQNYLE KKVEGHLQDE LKSNCANEVH 360 IEETINYPLS VRRTLLDSNE SLKKVDSFSR WITKALGEVD NLNMQSSPGI SWSTDECGHV 420 IDDTSLSPSL SQDQLYSIND FSPKWAYAGS DTEVLIIGSF LKSQPEVTTY NWSCMFGEVE 480 VPAEVVANGI LCCQAPPHKV GRVPFYVTCS NRLACSEVRE FDFREGYSSN VDYTDFFNSS 540 NDMLLHLRLD KFLSLKPVHP SNQAFEGDME KINLIFKLIS LREEEDYSSK EEKTVEMNIS 600 RHKVKEHQFH RQFKENLYSW LLHKVTESGK GPNVLDKDGQ GVLHLAAVLG YYWAITPILI 660 AGVNVNFRDV NGWTALHWAA SCGRERTVAV LVSMGADCGA LTDPSPEFPS GRTAADLASS 720 NGHKGISGFL AESSLTSHLE SLTVDDKQKG GQQEISGTKA VQTVSERTAT PVVYNDMPDG 780 LCLKDSLTAV RNATQAADRI HQVFRMQSFQ RKQLTQYEDD EFGLSDQRAL SLLASKVCKS 840 GQRDGLVNVA ATQIQKKFRG WKKRKEFLII RERIVKIQAH VRGHQVRKQY KTIIWSVGIL 900 EKVILRWRRK GSGLRGFRPD TLNKAPSQQS DSLKEDDYDY LKEGRKQKEE KIEKALSRVK 960 SMVQYPEARA QYRRVLNVVE DFRQKKDSNM GLISSEETVD GVEDLIDIDM LLDDDNFIPI 1020 AFD |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5hry_A | 9e-12 | 615 | 732 | 46 | 157 | ank3C2_1 |
| 5hry_B | 9e-12 | 615 | 732 | 46 | 157 | ank3C2_1 |
| 5hry_C | 9e-12 | 615 | 732 | 46 | 157 | ank3C2_1 |
| 5hry_D | 9e-12 | 615 | 732 | 46 | 157 | ank3C2_1 |
| 5hry_E | 9e-12 | 615 | 732 | 46 | 157 | ank3C2_1 |
| 5hry_F | 9e-12 | 615 | 732 | 46 | 157 | ank3C2_1 |
| 5hry_G | 9e-12 | 615 | 732 | 46 | 157 | ank3C2_1 |
| 5hry_H | 9e-12 | 615 | 732 | 46 | 157 | ank3C2_1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
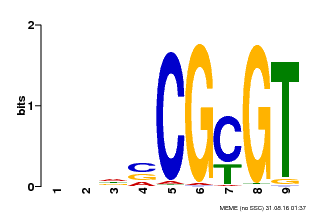
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_004485582.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC187465 | 0.0 | AC187465.3 Medicago truncatula chromosome 2 BAC clone mth2-69g14, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004485582.1 | 0.0 | calmodulin-binding transcription activator 2-like isoform X1 | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A1S2X9F7 | 0.0 | A0A1S2X9F7_CICAR; calmodulin-binding transcription activator 2-like isoform X1 | ||||
| STRING | XP_004485582.1 | 0.0 | (Cicer arietinum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF7406 | 26 | 42 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



