 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | CA08g15740 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Solanoideae; Capsiceae; Capsicum
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1019aa MW: 114798 Da PI: 6.138 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 178.8 | 6.7e-56 | 20 | 136 | 3 | 118 |
CG-1 3 ke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcyw 100
+e ++rwl++ ei++iL n++k++lt e++ rp sgs++L++rk++ryfrkDG++w+kkkdgktv+E+hekLKvg+++vl+cyYah+ee+ +fqrr+yw
CA08g15740 20 SEvQHRWLRPAEICEILRNYRKFHLTPEAPYRPVSGSVFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHEKLKVGSIDVLHCYYAHGEEDDNFQRRSYW 118
5559*********************************************************************************************** PP
CG-1 101 lLeeelekivlvhylevk 118
+Le++l +iv+vhylevk
CA08g15740 119 MLEQDLMHIVFVHYLEVK 136
***************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 80.591 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 3.1E-79 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 4.2E-49 | 22 | 135 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 2.9E-6 | 469 | 555 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 5.0E-7 | 470 | 555 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 2.33E-17 | 470 | 556 | IPR014756 | Immunoglobulin E-set |
| SuperFamily | SSF48403 | 3.58E-15 | 659 | 764 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 8.4E-15 | 666 | 768 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.24E-12 | 668 | 762 | No hit | No description |
| PROSITE profile | PS50297 | 16.335 | 670 | 774 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 9.484 | 703 | 735 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.0036 | 703 | 732 | IPR002110 | Ankyrin repeat |
| SMART | SM00015 | 5.8 | 877 | 899 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.023 | 878 | 907 | IPR000048 | IQ motif, EF-hand binding site |
| SuperFamily | SSF52540 | 2.02E-7 | 878 | 928 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| Pfam | PF00612 | 0.0029 | 879 | 898 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0012 | 900 | 922 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.542 | 901 | 925 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 7.0E-5 | 903 | 922 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1019 aa Download sequence Send to blast |
MADCGSDPSG FRLDITQILS EVQHRWLRPA EICEILRNYR KFHLTPEAPY RPVSGSVFLF 60 DRKVLRYFRK DGHNWRKKKD GKTVKEAHEK LKVGSIDVLH CYYAHGEEDD NFQRRSYWML 120 EQDLMHIVFV HYLEVKGNKV NVGYIRSVKF AHSNYQNECS LSDSMPTGHK KLASANADSA 180 SLASTLTEAH EEAESEDNHH ACSRFHSYPD RASGMDSHLV ESRDTICSSY GSPQSSVEYT 240 SLPSIDGAGK CDLGNFASGP QRTVDLGYGF PVSQHCSNGE MVGQDDFKNN LSVHENWQCS 300 FGDSPLQFHG QNVNQDLIAD LSYGLGNSFQ NRSLPSDLLS VRGQSYLYPD AQEGQLTQLD 360 LQYLNSLLEV QGDMNQESNM DMIELGDYST VKQPQLSSVK MEEGLKKVDS FSRWVAKELE 420 DVEELHMQPN NRISWNAIDT EEEDSYLPNR LHMDSDSLNP SLSQEQVFSI IDFSPNWAYS 480 NFETKVLITG RFLKSEGELV EYKWSCMFGE VEVPAEVLAD GVLRCHAPPH KPGVLPFYVT 540 CSNRLACSEV REFEYRFGPF QEFGAANVST TEMHLLERIE NLLSLGPVSN CRISDTMEAA 600 KDKQSTVNKI IFMMEEENQQ MIERASDYDT SQCRVKEDLF LESKLKQNFY AWLIHQVTDD 660 GRGRTLLDDE GQGILHLVAA LGYDWALKPI LASGVSVDFR DINGWTALHW AAFYGREKTV 720 VGLVSLGASP GALTDPSAEF PLARTPADLA SANGHKGISG FLAESSLTTH LTKLTVSDAK 780 EELASEGCEA KVGETVTERV AVTTTGNDVP DVLSLKDSLA AIRNATQAAA RIHQIFRVQS 840 FQRKQIIEHS DDELSSDENA LSILASKSCK LGQNNGIAHA AAIQIQKKFR GWNKRKEFLL 900 IRQKIVKIQA HIRGHQVRKK YKPIIWSVGI LEKVILRWRR KRSGLRGFKS EEVMNKPSTQ 960 DDSLPEDDYD FLKEGRKQTE VRMQKALARV KSMTQYPEGR AQYRRLLTAA EGLREVKV* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cxk_A | 1e-13 | 470 | 559 | 9 | 92 | calmodulin binding transcription activator 1 |
| 2cxk_B | 1e-13 | 470 | 559 | 9 | 92 | calmodulin binding transcription activator 1 |
| 2cxk_C | 1e-13 | 470 | 559 | 9 | 92 | calmodulin binding transcription activator 1 |
| 2cxk_D | 1e-13 | 470 | 559 | 9 | 92 | calmodulin binding transcription activator 1 |
| 2cxk_E | 1e-13 | 470 | 559 | 9 | 92 | calmodulin binding transcription activator 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
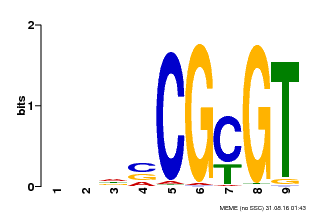
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | JN558810 | 0.0 | JN558810.1 Solanum lycopersicum calmodulin-binding transcription factor SR1L mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016539240.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 2 isoform X1 | ||||
| Refseq | XP_016539241.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 2 isoform X2 | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A1U8DYK6 | 0.0 | A0A1U8DYK6_CAPAN; calmodulin-binding transcription activator 2 isoform X2 | ||||
| TrEMBL | A0A1U8E7F6 | 0.0 | A0A1U8E7F6_CAPAN; Uncharacterized protein | ||||
| TrEMBL | A0A2G2Y333 | 0.0 | A0A2G2Y333_CAPAN; calmodulin-binding transcription activator 2 isoform X1 | ||||
| STRING | XP_009600617.1 | 0.0 | (Nicotiana tomentosiformis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA2230 | 21 | 46 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



